Tools, Publications, & Data
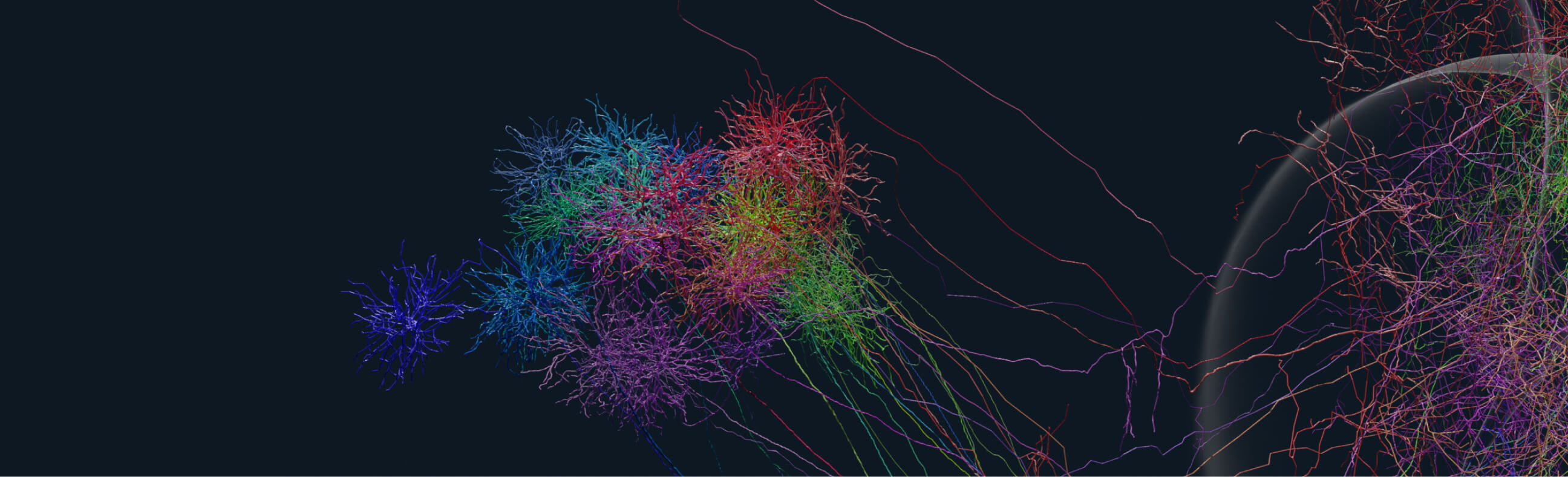



Public Code Ocean collections we have developed
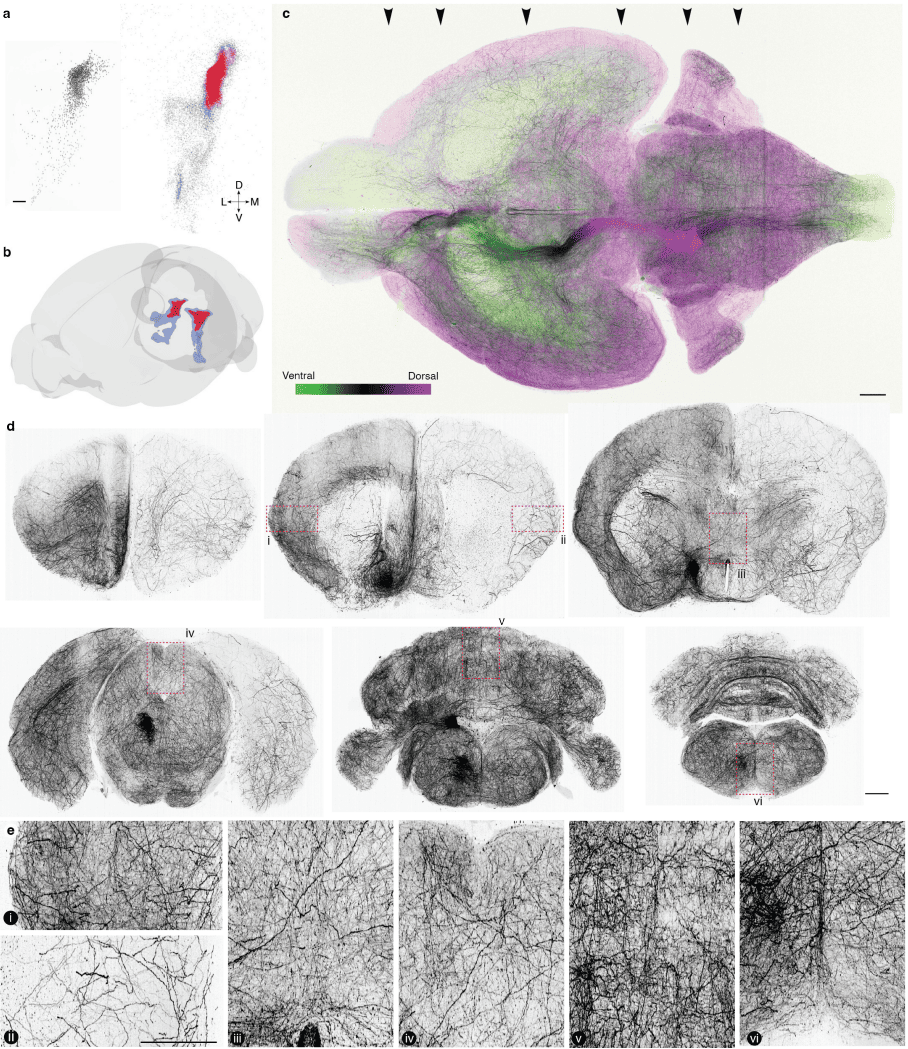
PREPRINT: We found a relationship between the morphologies, gene expression, and activity of LC-NE neurons in mice. These observations reveal a topographically organized structure and function of a neurotransmitter system and show that it contains learning signals for flexible behavior.
A collection of public Code Ocean capsules and data that accompany the paper "Topographic structure and function of locus coeruleus norepinephrine neurons"
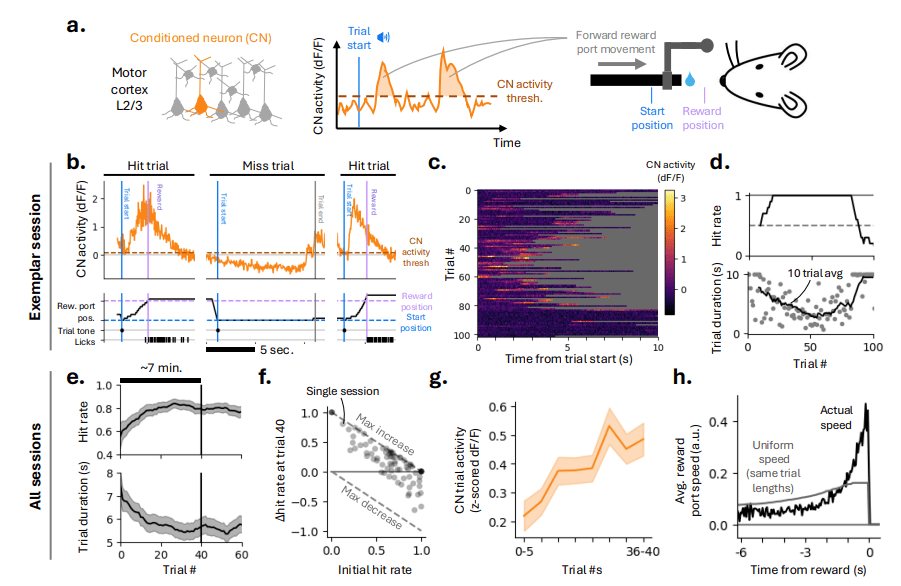
PREPRINT: We used two-photon photostimulation and calcium imaging to map learning-related changes in connectivity in layer 2/3 of mouse motor cortex. Our findings show how rapid learning can be achieved through structured changes in motor cortex connectivity.
Repository that hosts the SmartSPIM pipeline code.

This document details the steps necessary to assemble a Variable Difficulty Upright Mouse Running Wheel with Allen Institute Headframe Clamp.

This protocol describes the surgical procedure, instrumentation, and reagents necessary for the installation of a stabilizing headframe and cranial window over the visual cortex of an adult mouse. This protocol should be utilized for neural imaging and experimentation regarding the mouse visual cortex.

A Harp-compliant, 16-channel MOSFET Output board to control solenoid valves on closed loop with submillisecond precision.

This protocol details the acute craniotomy procedure to target locus coeruleus (LC) in vivo with a Neuropixels 2.0 single shank probe, and includes the skull thinning procedure to better expose regions of cortex for antidromic laser stimulation.
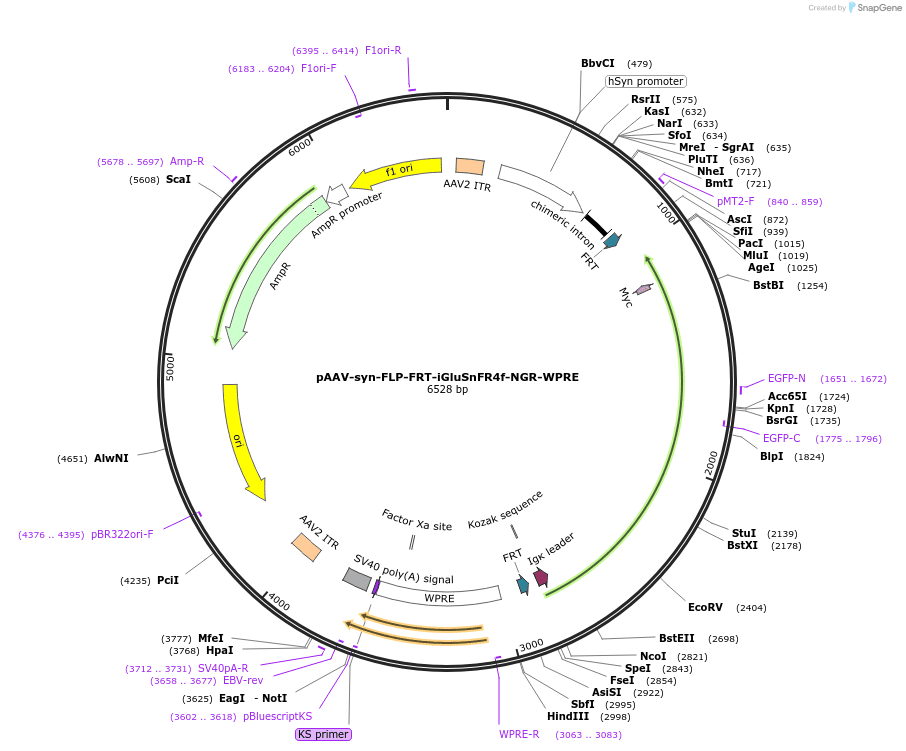
AAV-mediated expression of glutamate sensor with improved sensitivity and deactivation kinetics; NGR vector matches or outperforms PDGFR in all conditions tested.
.jpeg)
This LifeCanvas SmartBatch+ protocol is intended for whole mouse brain tissue clearing and incorporates electrophoretic delipidation. While this delipidation protocol is longer than other delipidation protocols we tested, it preserves the endogenous fluorescence of the tissue and clears the whole brain most effectively, demonstrating the morphology of fine cell structures deep in the brain.

This protocol discusses how to mount the sample holder into the microscope and the steps to acquire a full-brain volumetric image using the custom ExA-SPIM software.
Closed-loop behavior system for continuous, long-term (multi-week) experiments in freely-moving mice.
The Genetic Tools Atlas (GTA) is a searchable web tool representing information and data on enhancer-adeno-associated viruses (enhancer AAVs) and mouse transgenes developed and tested at the Allen Institute for Brain Science. The GTA offers a large genetic toolkit for selective gene expression in brain cell types of interest.
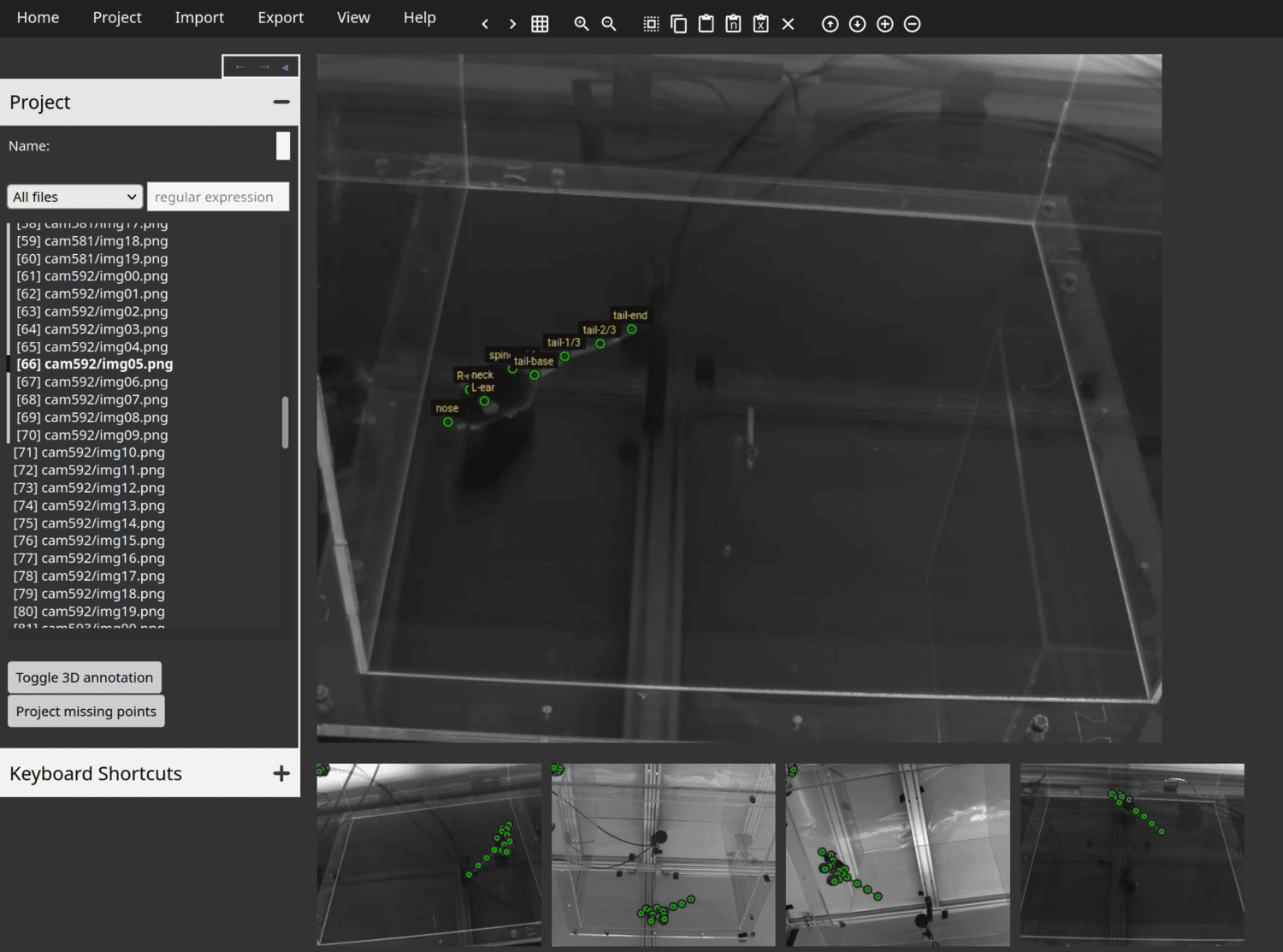
Anivia is a new web tool intended specifically for annotation of animal keypoints from images in both 2D and 3D.
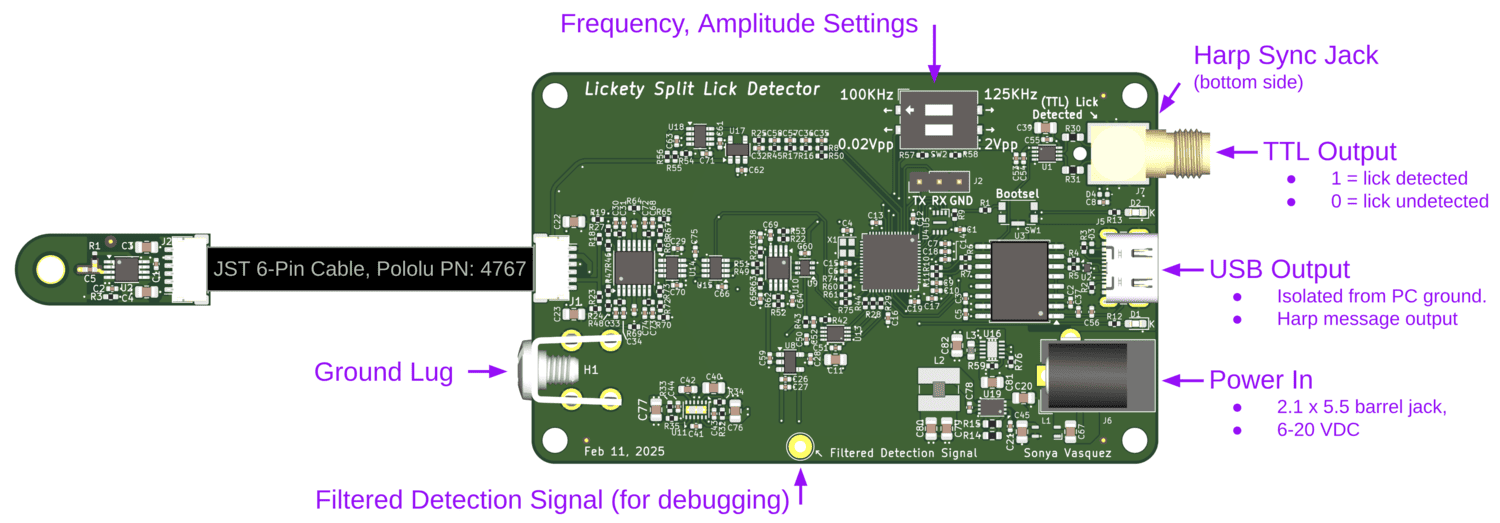

Several Harp devices:

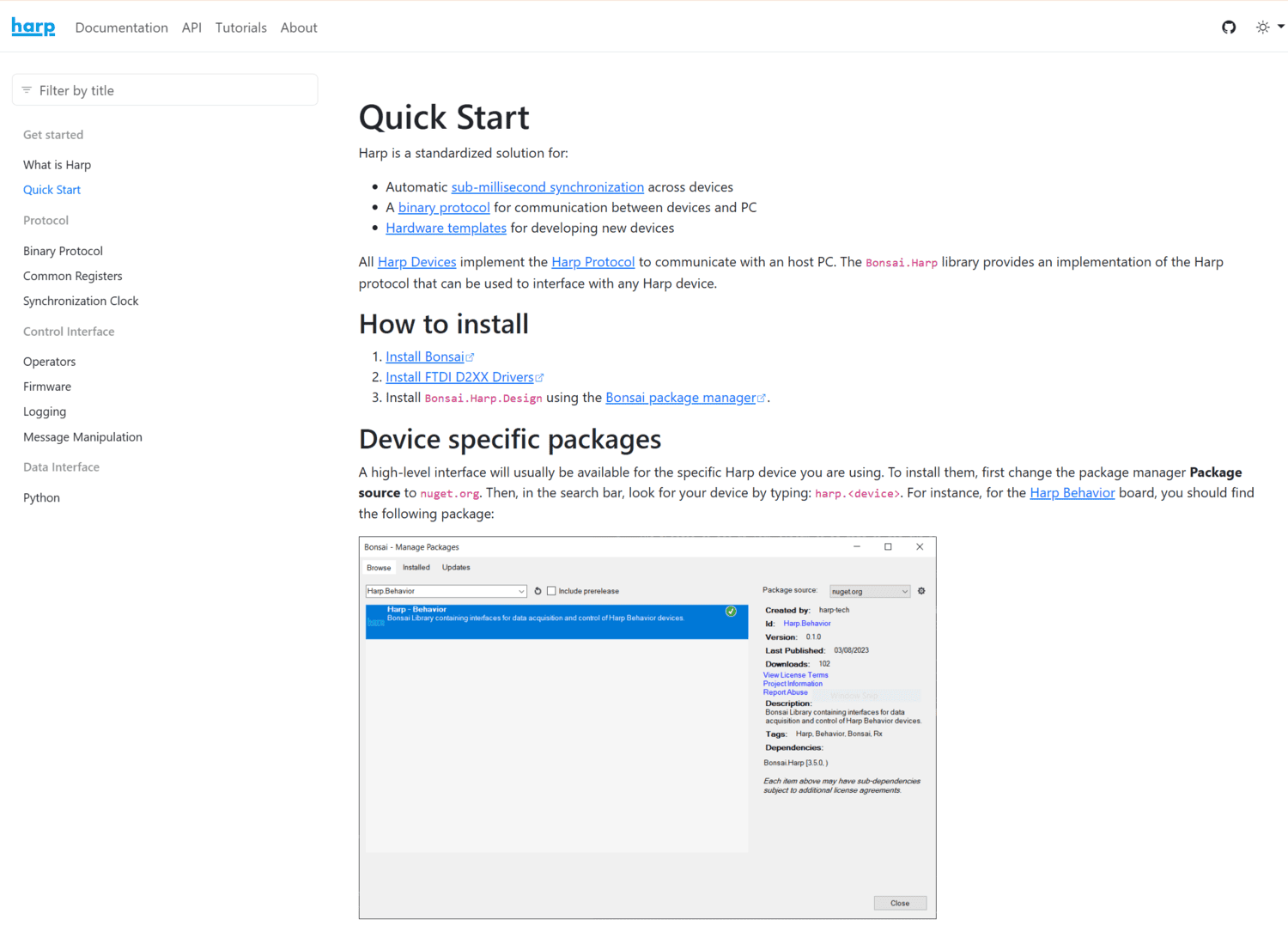
Harp is a standardized solution for:
All Harp Devices implement the Harp Protocol to communicate with an host PC. The Bonsai.Harp library provides an implementation of the Harp protocol that can be used to interface with any Harp device.
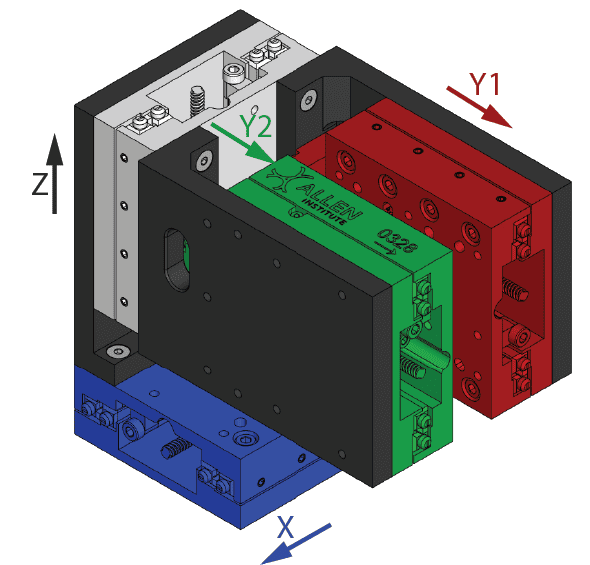
Software that controls the manipulator. The manipulator is composed of two main components:
A repository for calibration and configuration of the harp olfactometer device
A core problem in mice training is accurately keeping track of each mouse’s training stage and accurately setting the corresponding rig parameters. As the number of behavior studies, research assistants, and mice increase, manual tracking and parameter input is prone to human error. This library provides a flexible framework for defining mice curriculum enabling mouse training to be automated.
A repository for the VR Foraging task.
A Harp core implementation on the RP2040
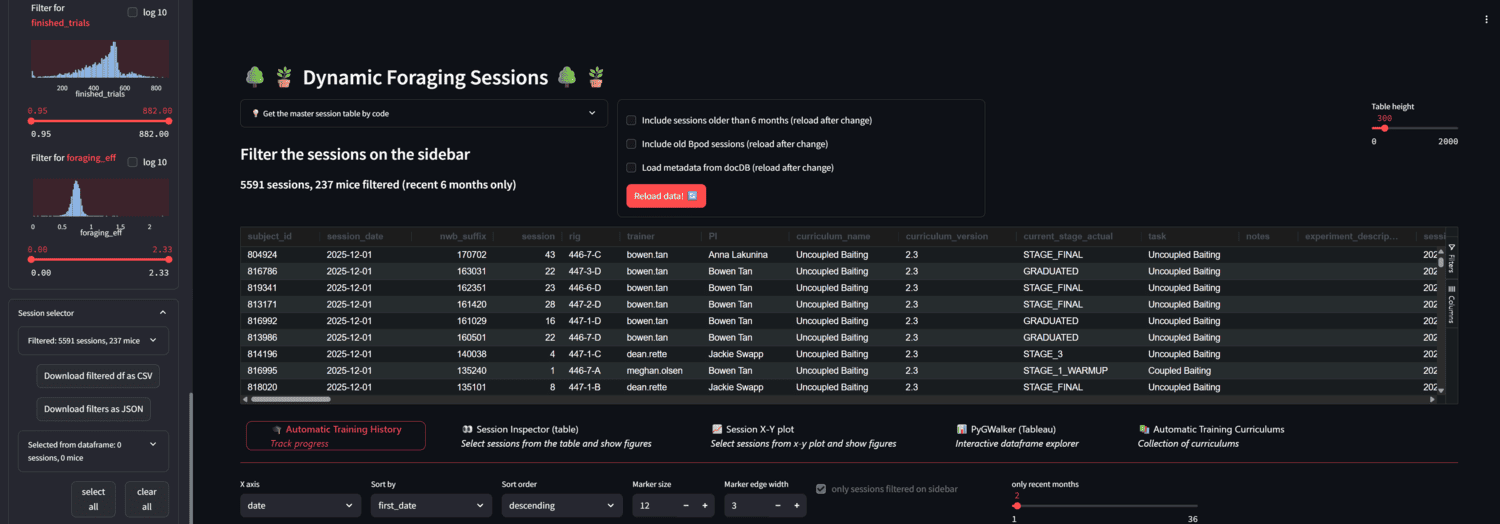
A streamlit app for browsing foraging behavior sessions in AIND.
A pubic collection accompanying "Efficient and reproducible pipelines for spike sorting large-scale electrophysiology data."

PREPRINT: We present an end-to-end spike sorting pipeline that leverages parallelization to scale to large datasets.

This modified LifeCanvas embedding protocol is intended to prepare cleared mouse brains for imaging on the SmartSPIM. It details mixing and degassing the agarose solution and the technical process of embedding a brain in the agarose solution.

This protocol describes the experimental setup and procedure for behavioral training of mice in a “dynamic foraging” task, wherein a thirsty mouse is trained to make decisions in a dynamic environment.


This protocol outlines the parameters, steps, and frequency of water calibration and spot checks to ensure that reward volumes remain consistent over time.
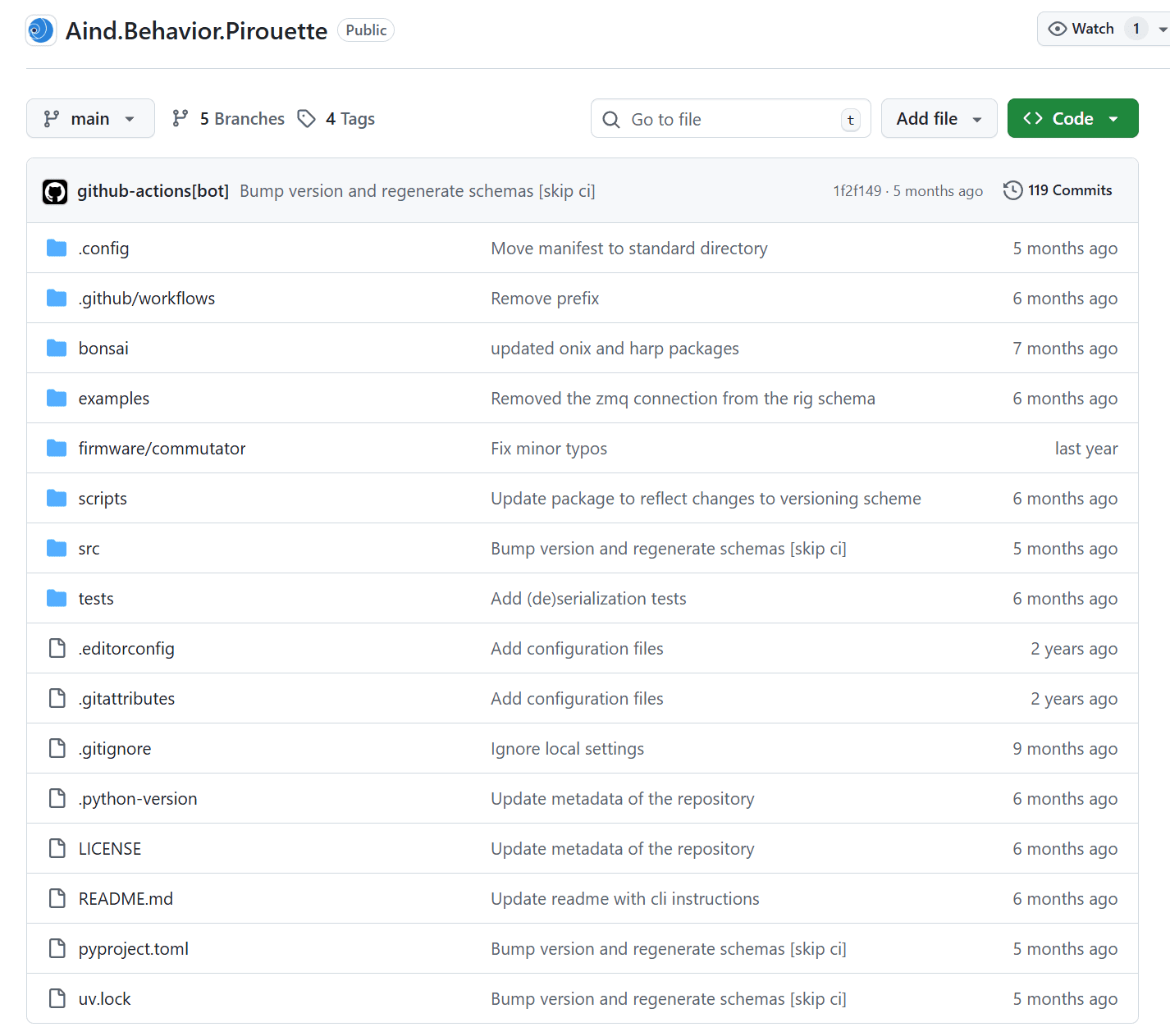
A repository with source code for long-term, freely moving, electrophysiology recordings in mice.
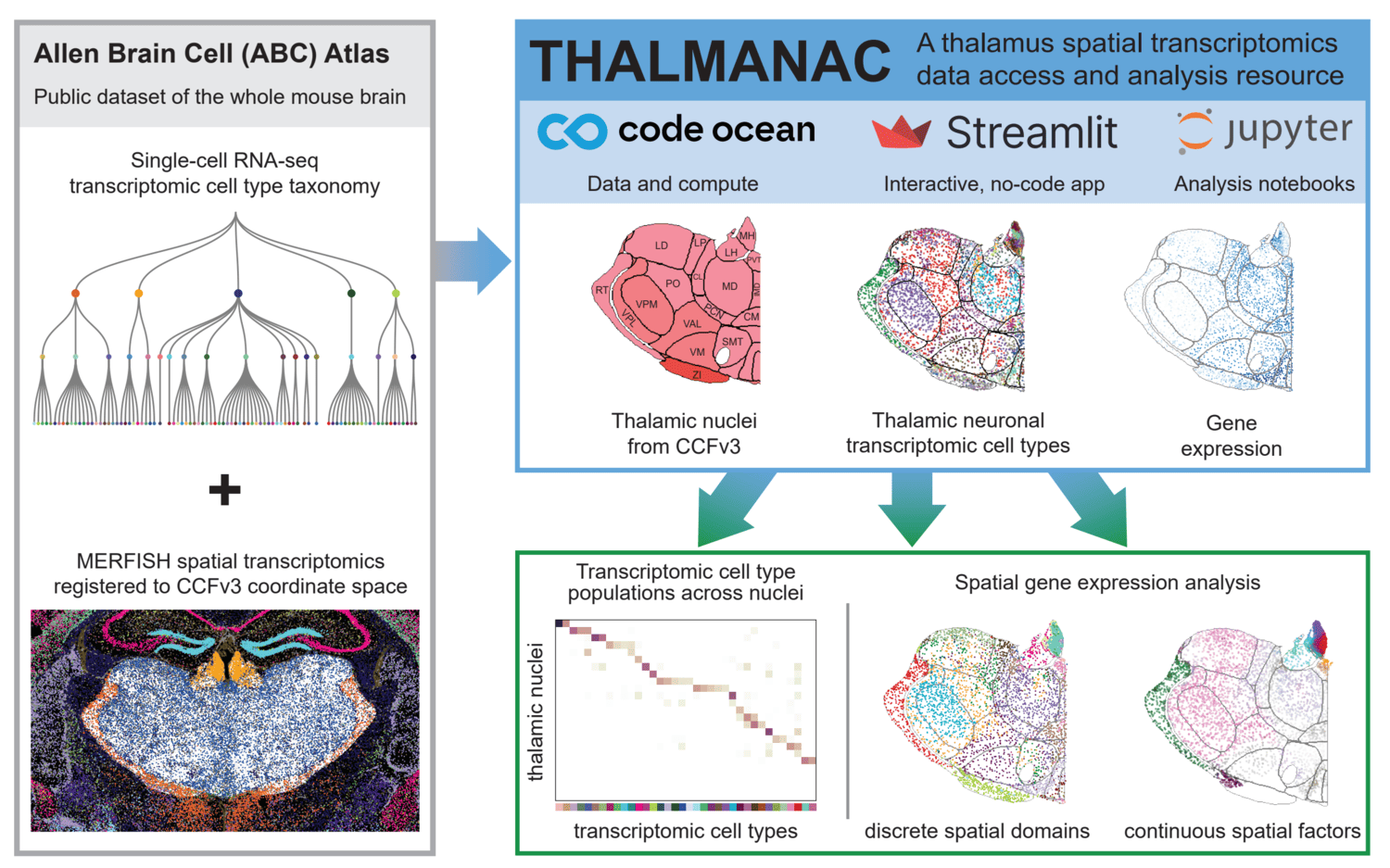
PREPRINT: We present the THALMANAC, (THALamus MERFISH ANalysis and ACcess) a Findable, Accessible, Interoperable, Reusable and Reproducible (FAIRR) resource for exploring and analyzing single-cell transcriptomic variation in the thalamus. The THALMANAC provides streamlined access to thalamic gene expression data registered to the common coordinate framework and tools for quantitative analysis and visualization of these data, all encapsulated in a reproducible, cloud computing platform.
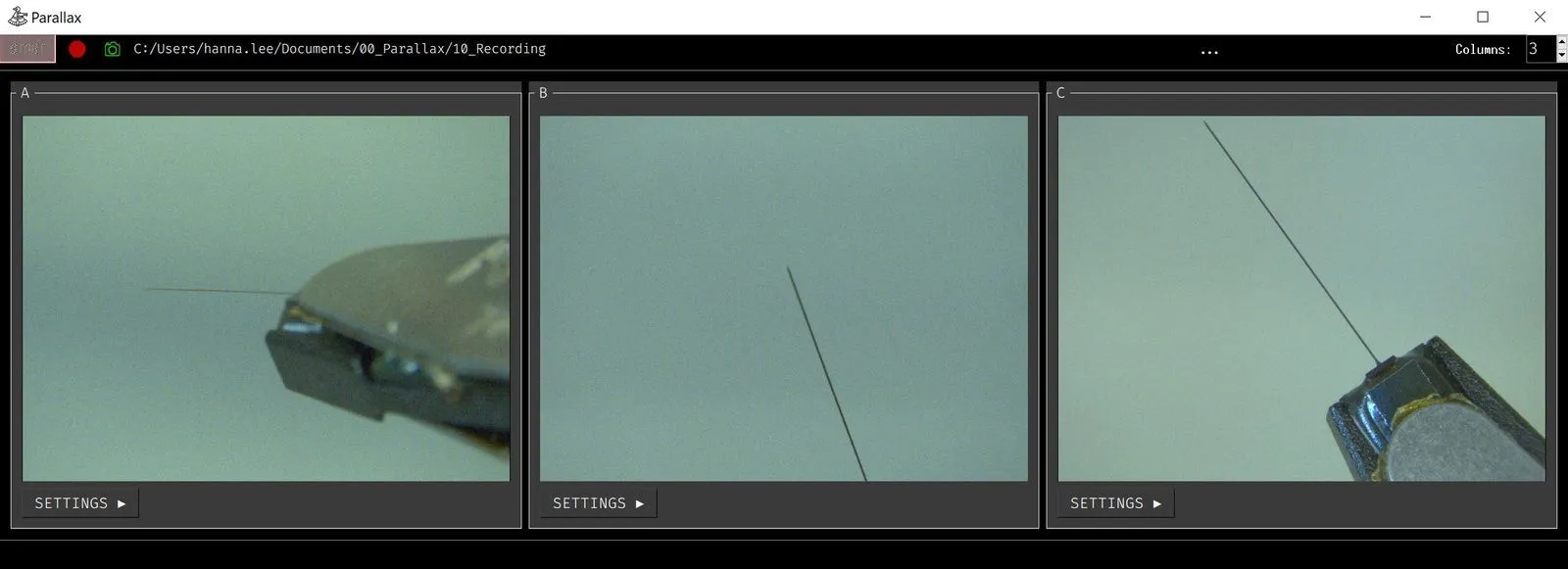
Parallax is a graphical user interface designed to streamline the process of setting up and performing acute in vivo electrophysiology experiments.

A flexible, scalable extracellular electrophysiology rig
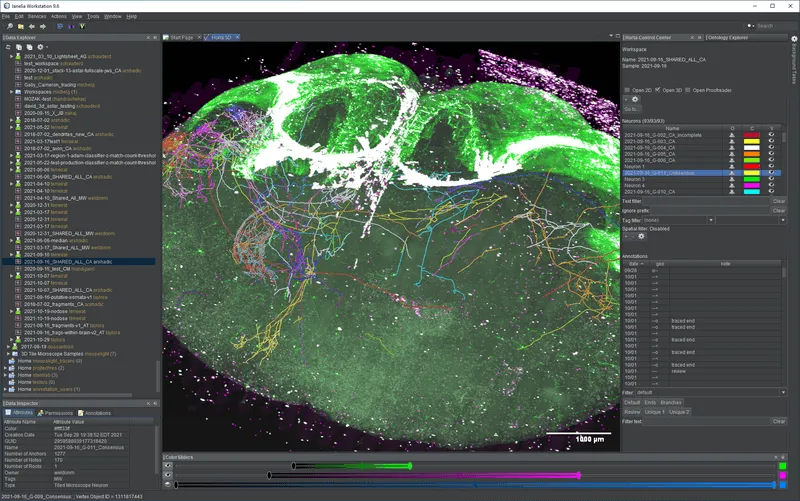
HortaCloud is a streaming 3D annotation platform for large microscopy data that runs entirely in the cloud. It is a free, open source research software tool, developed by Janelia Research Campus. The Allen Institute uses HortaCloud to proofread whole-brain images of neurons collected on various fluorescence microscopy instruments.
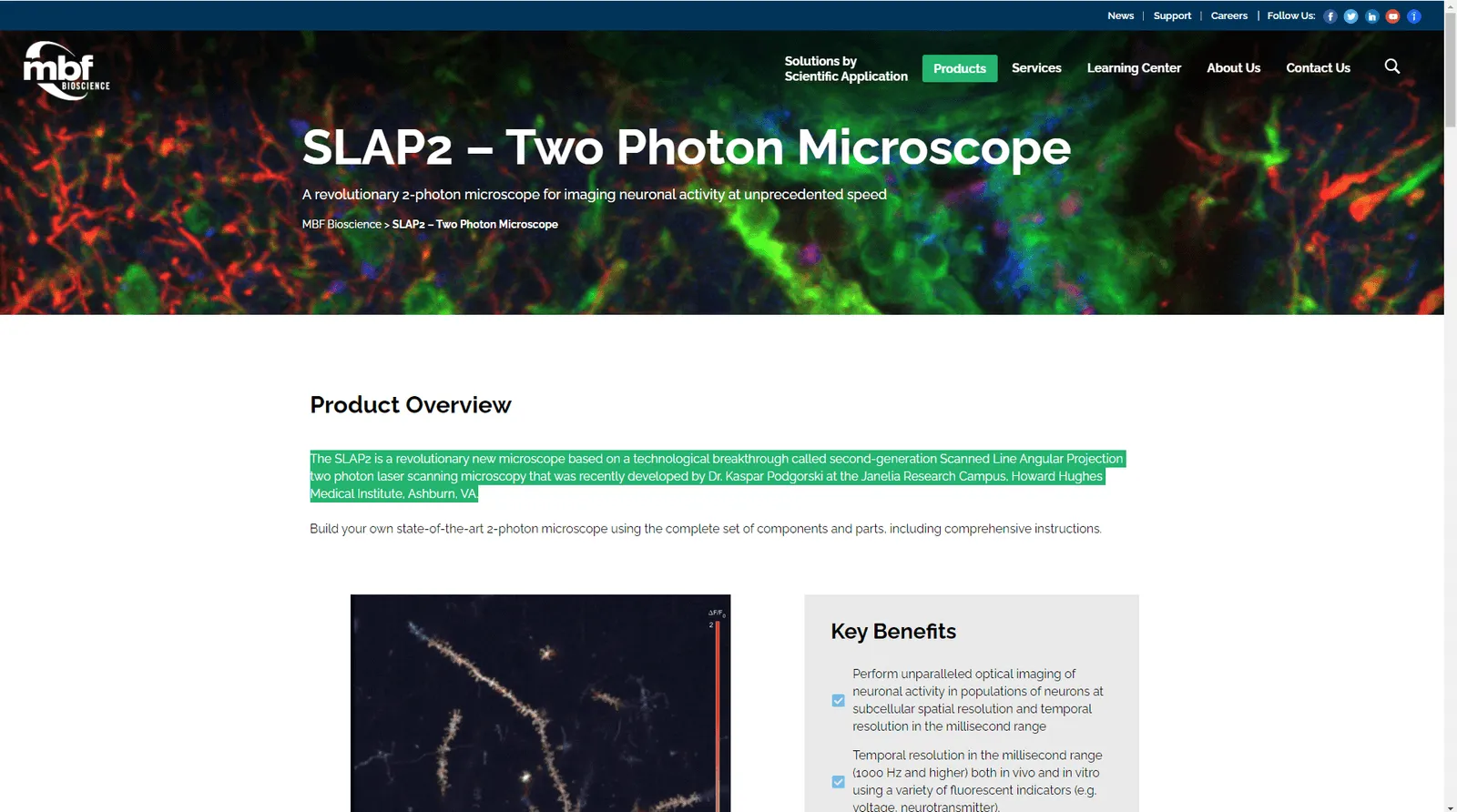
The SLAP2 is a revolutionary new microscope based on a technological breakthrough called second-generation Scanned Line Angular Projection two photon laser scanning microscopy that was recently developed by Dr. Kaspar Podgorski at the Janelia Research Campus, Howard Hughes Medical Institute, Ashburn, VA.
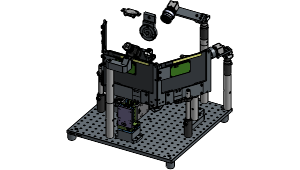

A Behavior platform designed for VR Foraging tasks, supporting multi-modal sensory inputs and integration with the Allen Institute headframe system.
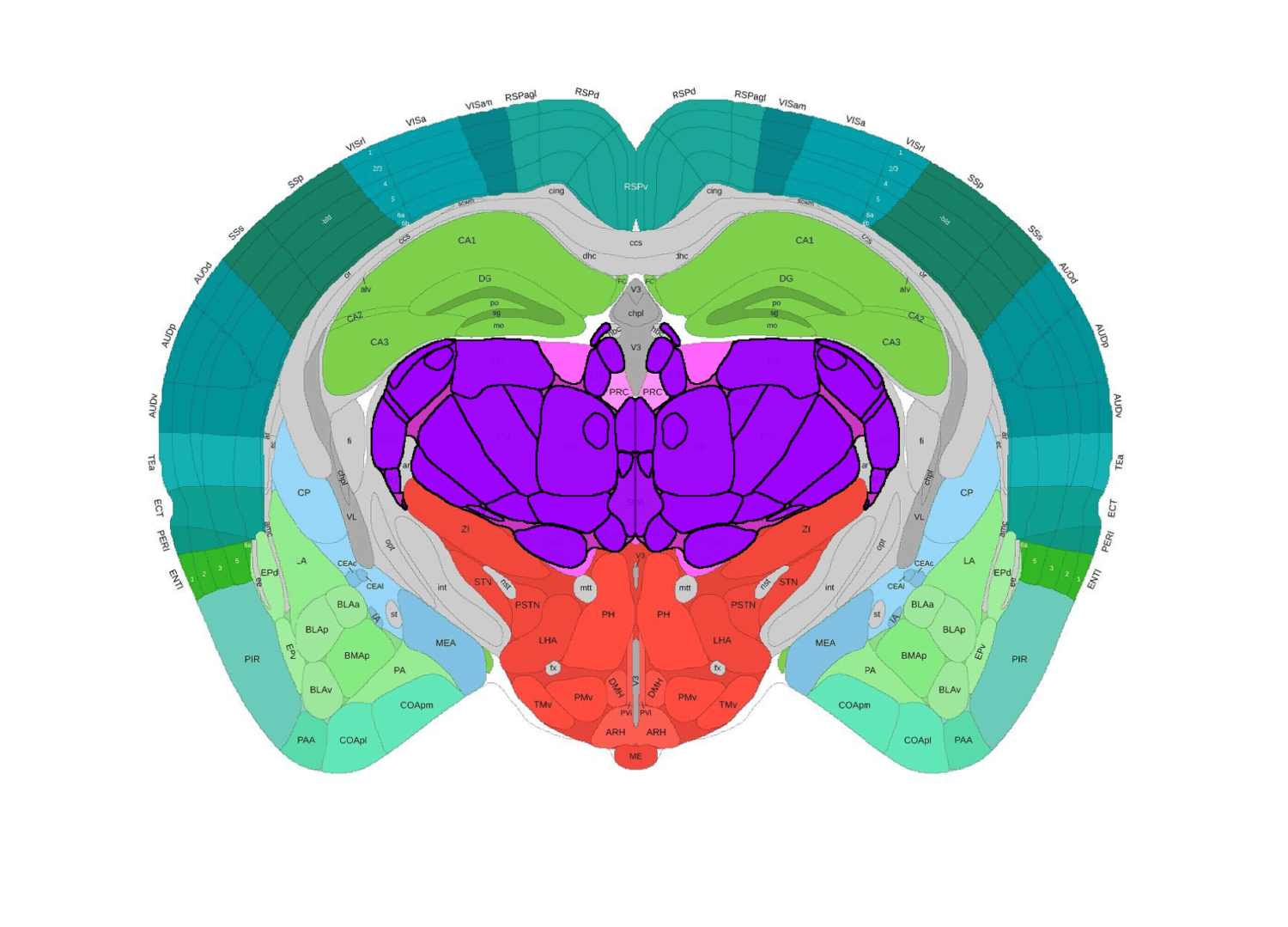
Registration of lightsheet whole brain images to the Allen CCF V3 Atlas

Code Ocean capsule and streamlit app for exploration, analysis, and easy access of thalamus MERFISH data. Jupyter notebooks in code contain walkthroughs of the major functionality, while those in code/figures directly reproduce manuscript figures.

This protocol describes the surgical procedure, instrumentation, and reagents necessary for the installation of a stabilizing headframe and cranial window over the cortex of an adult mouse. This protocol should be utilized for neural imaging and experimentation regarding the mouse cortex.
On this repository you can find documentation of the Harp protocol, what is a Harp device and the Harp Timestamp Bus. The Harp technology was originally thought at Champalimaud Foundation to suppress the lack of efficient binary protocols for scientific hardware communication.
Package for loading and interacting with AIND mesoscale anatomy whole-brain data.
Library for aind mri utilities.
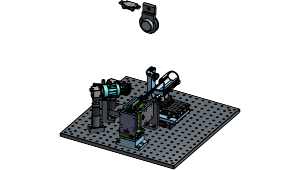

A behavior platform designed for Dynamic Foraging tasks, supporting multi-modal sensory inputs and integration with the Allen Institute headframe system.
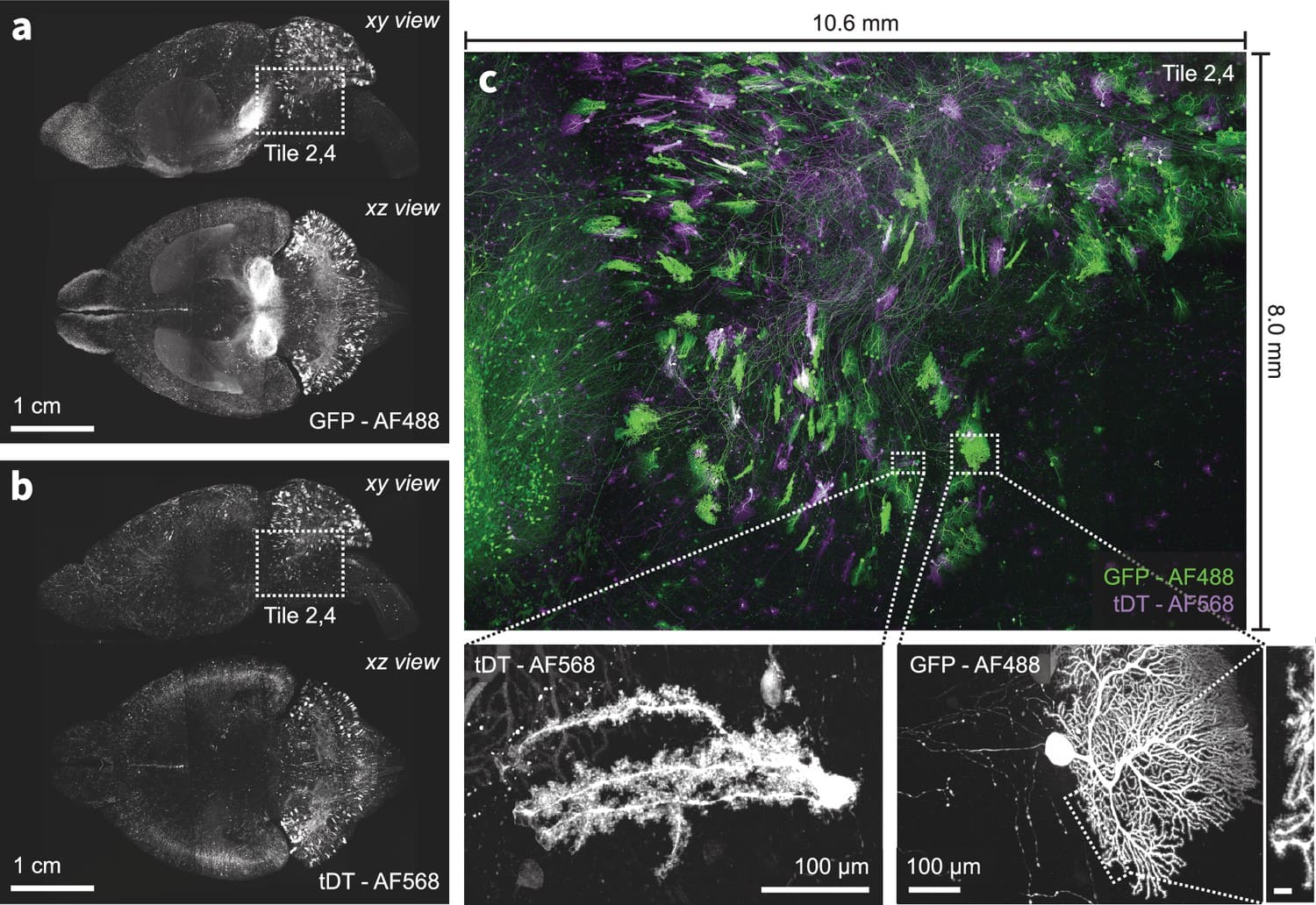
We present a new expansion-assisted selective plane illumination microscope, which, combined with new tissue clearing and expansion methods, the microscope allows imaging centimeter-scale samples with 250×250×750 nm optical resolution (4× expansion), including entire mouse brains, with high contrast and without sectioning.

BARseq is a high throughput in situ transcriptomics method utilizing designed barcodes incorporated in the amplification process for multiplexed gene detection.

Predictive processing theorizes that the brain continuously predicts sensory inputs, refining neuronal responses by minimizing prediction errors. Our collaborative experiments test these mechanisms using mismatch stimuli in mice and primates. The resulting datasets are shared via the OpenScope program and from collaborating laboratories, fostering community analysis and iterative refinement of predictive processing models.
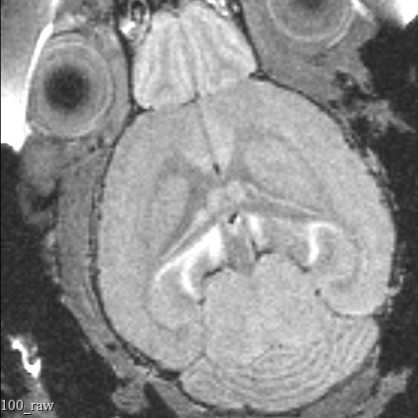
This protocol describes the process of obtaining pre-insertion MRI images using the University of Washington 14T vertical bore Bruker MRI. These structural scans allow for precisely targeting acute electrophysiology recordings in head-fixed mice beyond what is possible using skull landmarks alone.

The OpenScope program publishes on an annual basis a detailed public report that summarizes all of our activities within the year. These reports highlight all developments of the year: new selected projects, completed projects, publications and technical developments of the platform.
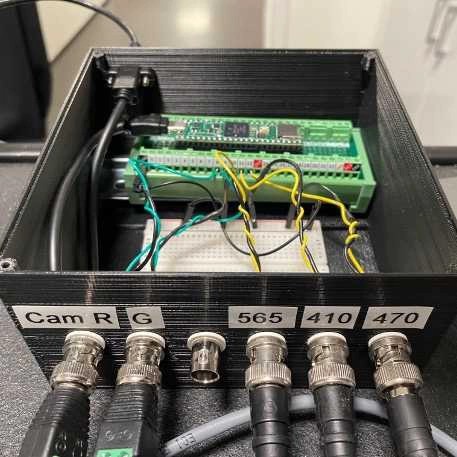

Frame-projected independent fiber photometry (FIP) is a method to measure fluorescent sensor signals through optical fibers implanted in living animals, using a camera to record video of the fiber faces. To precisely control timings of excitation and camera frame acquisition for FIP, and of other external stimulation apparatus (e.g. LEDs for photo-stimulation), we use a Teensy 4.1 microcontroller, which generates voltage pulses without relying on continuous communication with the operating system of the experimental computer for timing. This protocol is for the 3 excitation LED + 2 collection CMOS cameras design.

A library for managing data contracts and quality control in behavioral datasets.

This protocol describes the steps for preparing 20X stock and experimental solution of Lipopolysaccharide (LPS) intended for intraperitoneal injections in mice at a concentration of 0.25 mg/mL.

This protocol describes Vascular Access Button (VAB) catheter blood collection intended for later assessment of immune responses across time to different IP injected substances in awake mice. This protocol is preceded by “Mouse Catheter Maintenance” protocol.

This protocol describes blood preparation steps required for analysis of cytokines using an Olink assay. This protocol is applied to samples collected during the protocol 'Temporal Assessment of Immune Response.'

This protocol describes how to perform catheter maintenance to preserve mouse catheter patency after Vascular Access Button (VAB) implant.
.jpg)
This is a step-by-step protocol to build a modified FIP (Frame-projected Independent Fiber Photometry) system. FIP was first implemented and reported by Kim et al. (2016). Their protocol is available here.
We have modified the previous design so that it can be built primarily with Thorlabs products (some products from other manufacturers are still required), which are off-the-shelf and easy to obtain with a reasonable lead time. The system described in this protocol is designed to record signals from 1) GFP-based sensors, 2) mApple-based sensors, and 3) isosbestic signals from up to 4-9 sites depending on fiber bundle selection (Fig. 1 and 2). When applied to measurements of other fluorescent protein-based sensors or spectrally shifted sensors, the selection of optical filters and/or excitation light sources should be modified accordingly. Also, the selection of the fiber patch-cable should be based on the number of simultaneously recorded sites and the diameter/NA of the fiber implants.

This protocol describes the surgical procedure, instrumentation, and reagents necessary for performing stereotactic injections and securing a headframe to an adult mouse brain for in-vivo recording procedures.

PREPRINT: Identifying the input-output operations of neurons requires measurements of synaptic transmission simultaneously at many of a neuron’s thousands of inputs in the intact brain. To facilitate this goal, we engineered and screened 3365 variants of the fluorescent protein glutamate indicator iGluSnFR3 in neuron culture, and selected variants in the mouse visual cortex. Two variants have high sensitivity, fast activation (< 2 ms) and deactivation times tailored for recording large populations of synapses (iGluSnFR4s, 153 ms) or rapid dynamics (iGluSnFR4f, 26 ms). By imaging action-potential evoked signals on axons and visually-evoked signals on dendritic spines, we show that iGluSnFR4s/4f primarily detect local synaptic glutamate with single-vesicle sensitivity. The indicators detect a wide range of naturalistic synaptic transmission, including in the vibrissal cortex layer 4 and in hippocampal CA1 dendrites. iGluSnFR4 increases the sensitivity and scale (4s) or speed (4f) of tracking information flow in neural networks in vivo.
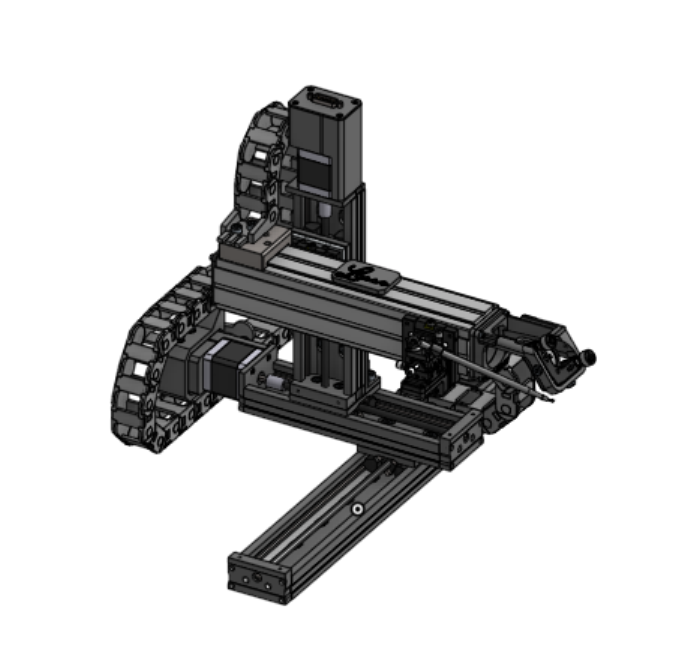

XYZ sample collection platform with loop collection and integrated optical system
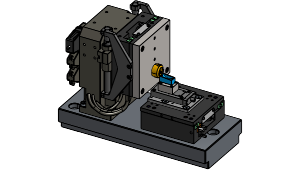

Microtome with slice thickness control and cutting actuation realized by high-precision linear stages

A reproducible system neuroscience notebook collection to facilitate data sharing and collaborative reuse with open science datasets

PREPRINT: Here, we evaluate the ability of the recently developed calcium indicator GCaMP8 to reveal neuronal spiking, and we investigate how existing spike inference methods (CASCADE, OASIS, MLSpike) should be adapted for optimal performance.
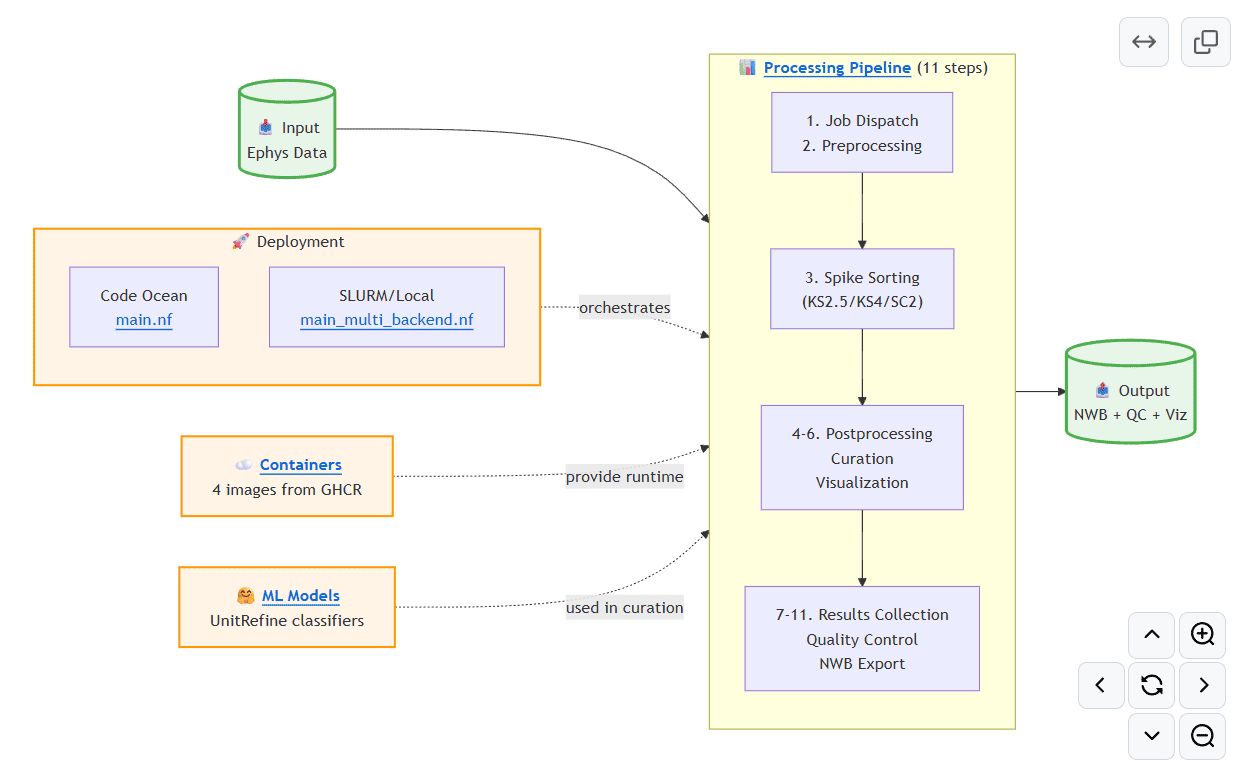
AIND pipeline for processing extracellular electrophysiology data
This protocol describes the surgical procedure, instrumentation, and reagents necessary for implanting optic fiber probes involving injection(s) and headpost into an adult mouse brain for in-vivo fiber photometry and/or optogenetic manipulation procedures.
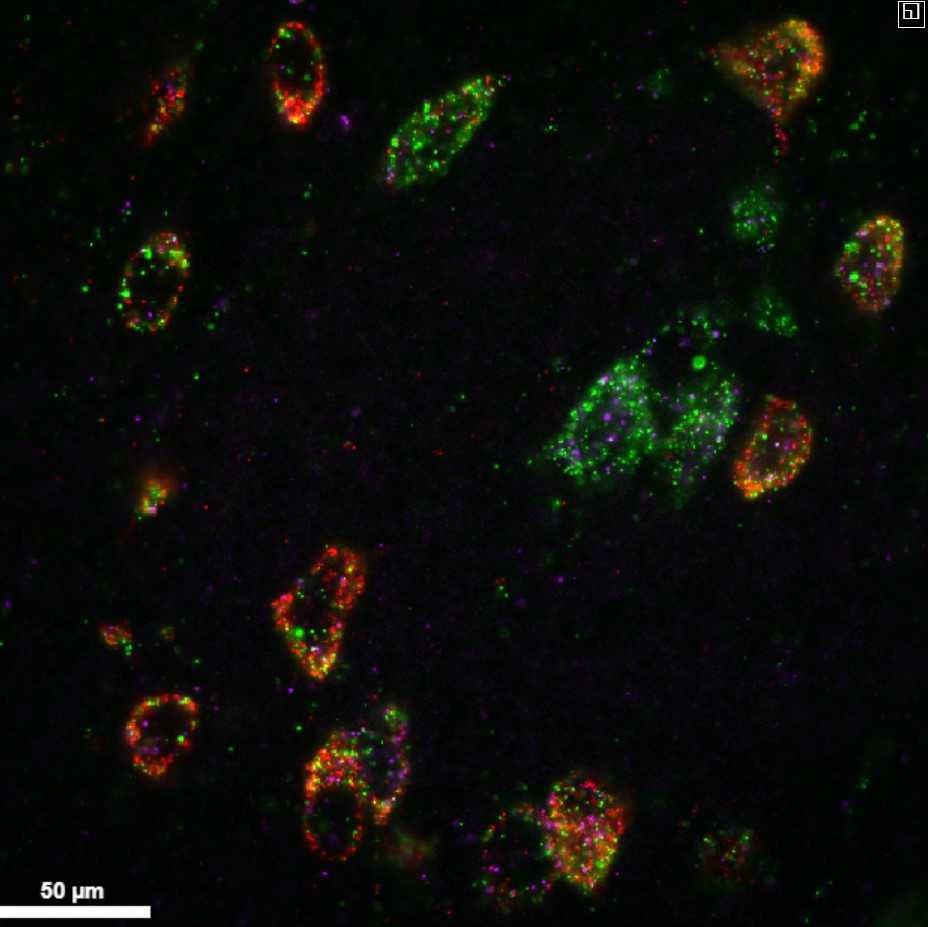
This protocol describes a method that utilizes expansion microscopy to perform multiple rounds of multiplexed fluorescent in situ hybridization (mFISH) on thick slices of brain tissue.

This protocol is modified from the Index Matching protocol found in the LifeCanvas Technologies Full Active Pipeline Protocol.
PREPRINT: Neuropixels Opto probes represent an unprecedented tool for recording, identifying, and manipulating neuronal populations by combining high-resolution extracellular electrophysiology with optogenetics.

This protocol details index matching for delipidated whole mouse brain samples in ethyl cinnamate solution to prepare for lightsheet imaging.
.jpeg)
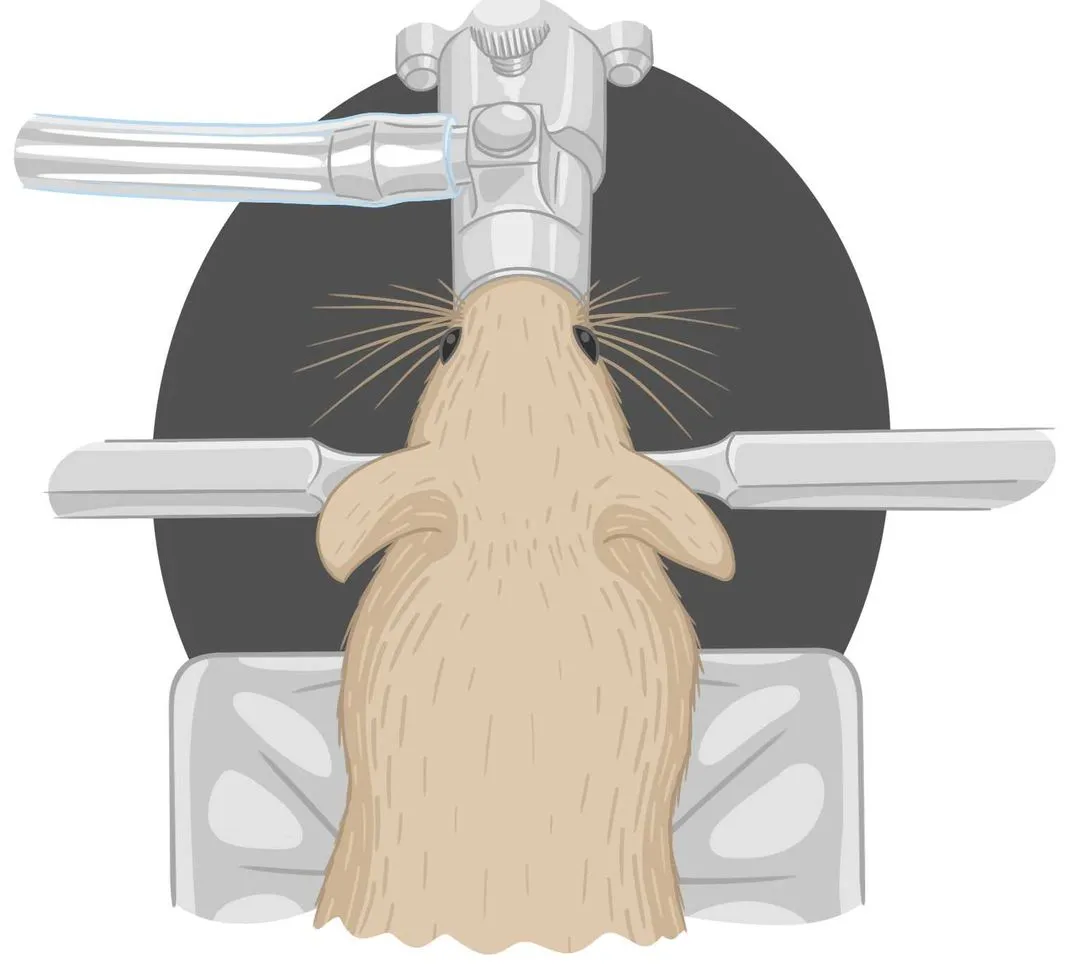
This protocol describes the typical workflow for the habituation of a mouse for awake behavioral experimentation that utilizes tubes to hold mice during head fixation.
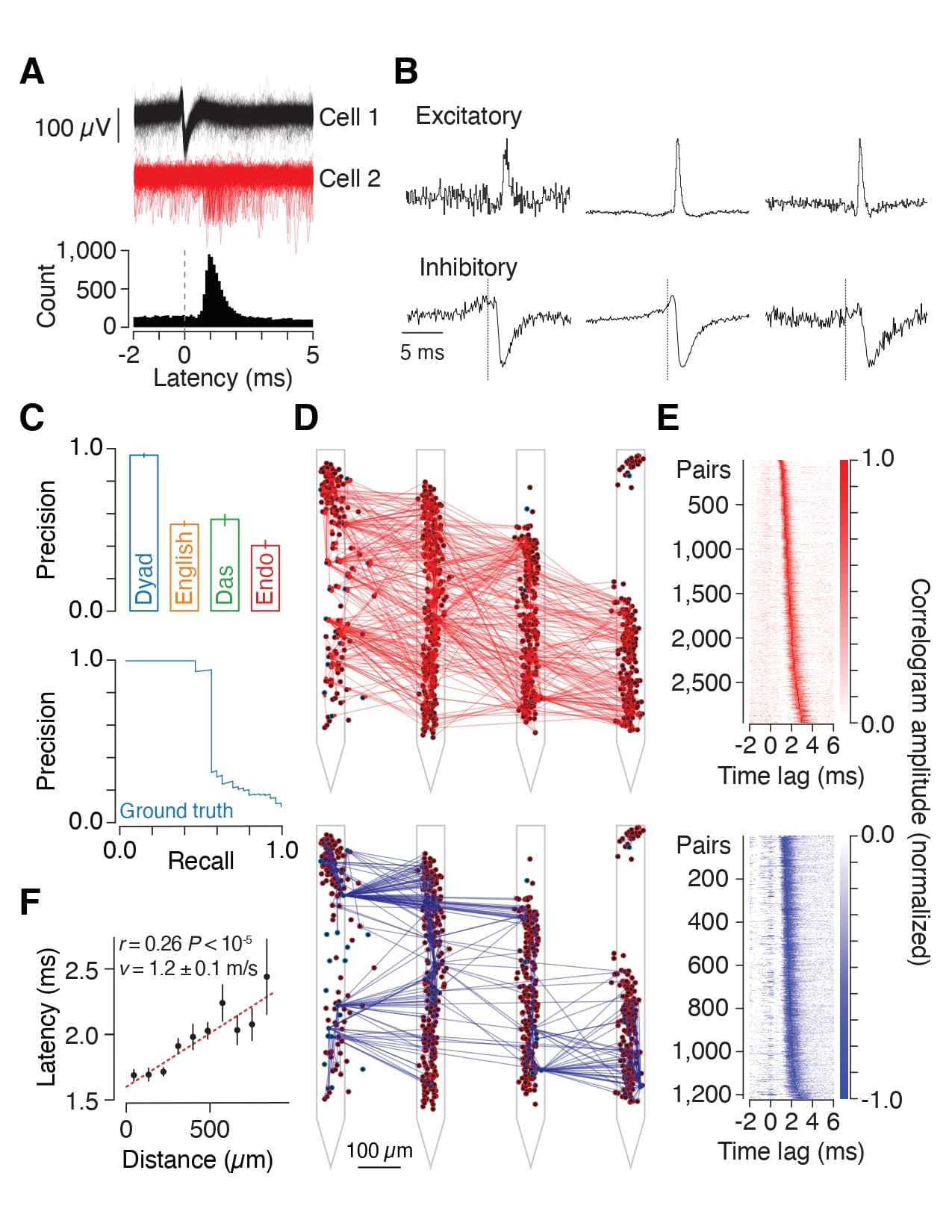
PREPRINT: We have studied how ongoing experience reorganizes the synaptic connectivity of neurons in the olfactory cortex of the mouse. This revealed that excitatory olfactory neurons with similar odor tuning are more likely to be connected, and inhibitory neurons that responded differentially to repeated encounters with a set of odorants exhibited a high degree of both incoming and outgoing synaptic connections within the cortical network.
.png)
PREPRINT: Predictive coding (PC) hypothesizes that the brain computes internal models of predicted events and that unpredicted stimuli are signaled with prediction errors that feed forward. We tested this hypothesis using a visual oddball task. A repetitive sequence interrupted by a novel stimulus is a “local” oddball. “Global” oddballs defy predictions while repeating the local context, thereby dissociating genuine prediction errors from adaptation-related responses. We recorded neuronal spiking activity across the visual hierarchy in mice and monkeys viewing these oddballs. Local oddball responses largely followed PC: they were robust, emerged early in layers 2/3, and fed forward. Global oddball responses challenged PC: they were weak, absent in most visual areas, more robust in prefrontal cortex, emerged in non-granular layers, and did not involve inhibitory interneurons relaying predictive suppression. Contrary to PC, genuine predictive coding does not emerge early in sensory processing, and is instead exclusive to more cognitive, higher-order areas.
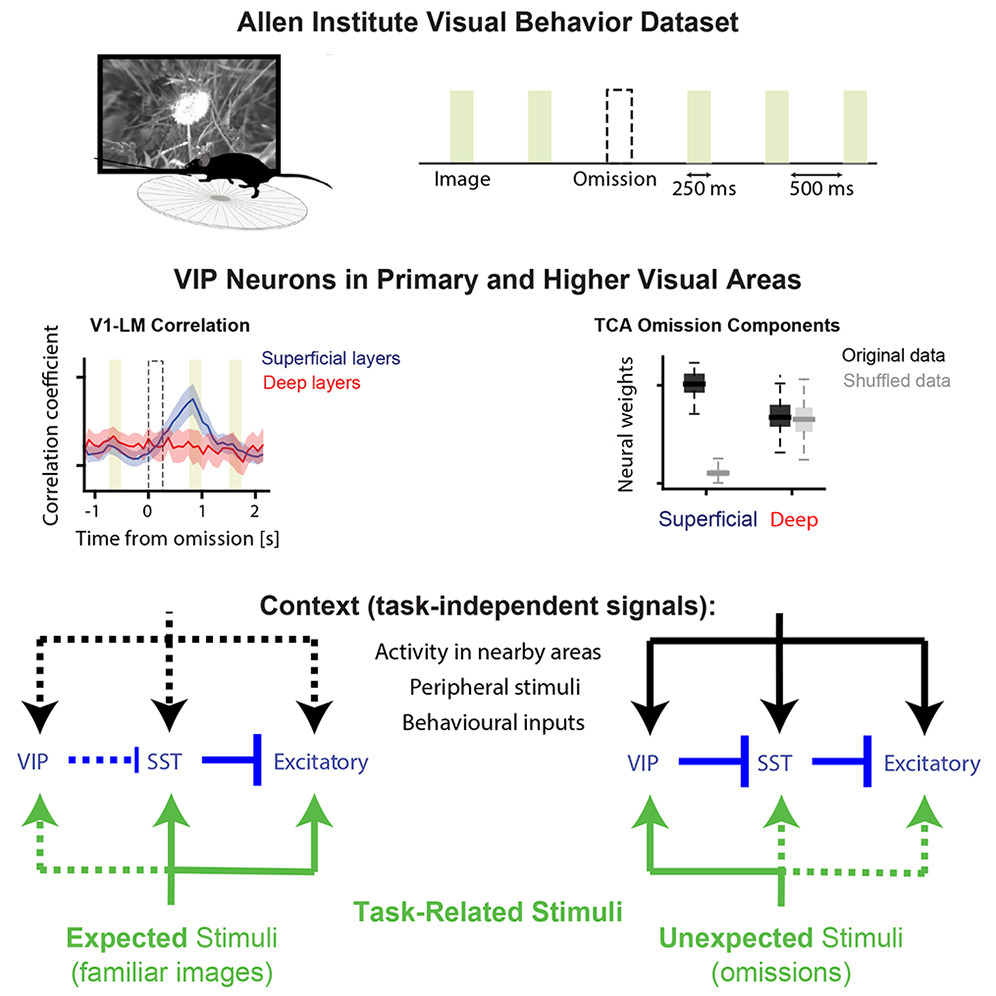
The visual cortex predicts incoming sensory stimuli through internal models that are updated following unexpected events. Cortical inhibitory neurons, particularly VIP interneurons, play a critical role in representing unexpected stimuli. Notably, this response is stimulus non-specific, raising the question of what information it conveys. Given their unique connectivity, we hypothesized that during unexpected stimuli, VIP neurons encode broad context signals, referred to here as task-independent information. To test this hypothesis, we analyzed the Allen Institute Visual Behavior dataset, in which mice viewed repeated familiar images and unexpected omissions of these images, while two-photon calcium imaging was performed from distinct cell types across primary and higher-order visual areas. Using dimensionality reduction methods, we found that, in contrast to image presentations, unexpected omissions trigger task-independent signaling in VIP and excitatory neurons. This signaling may facilitate the integration of contextual and sensory information, enabling updated predictions.
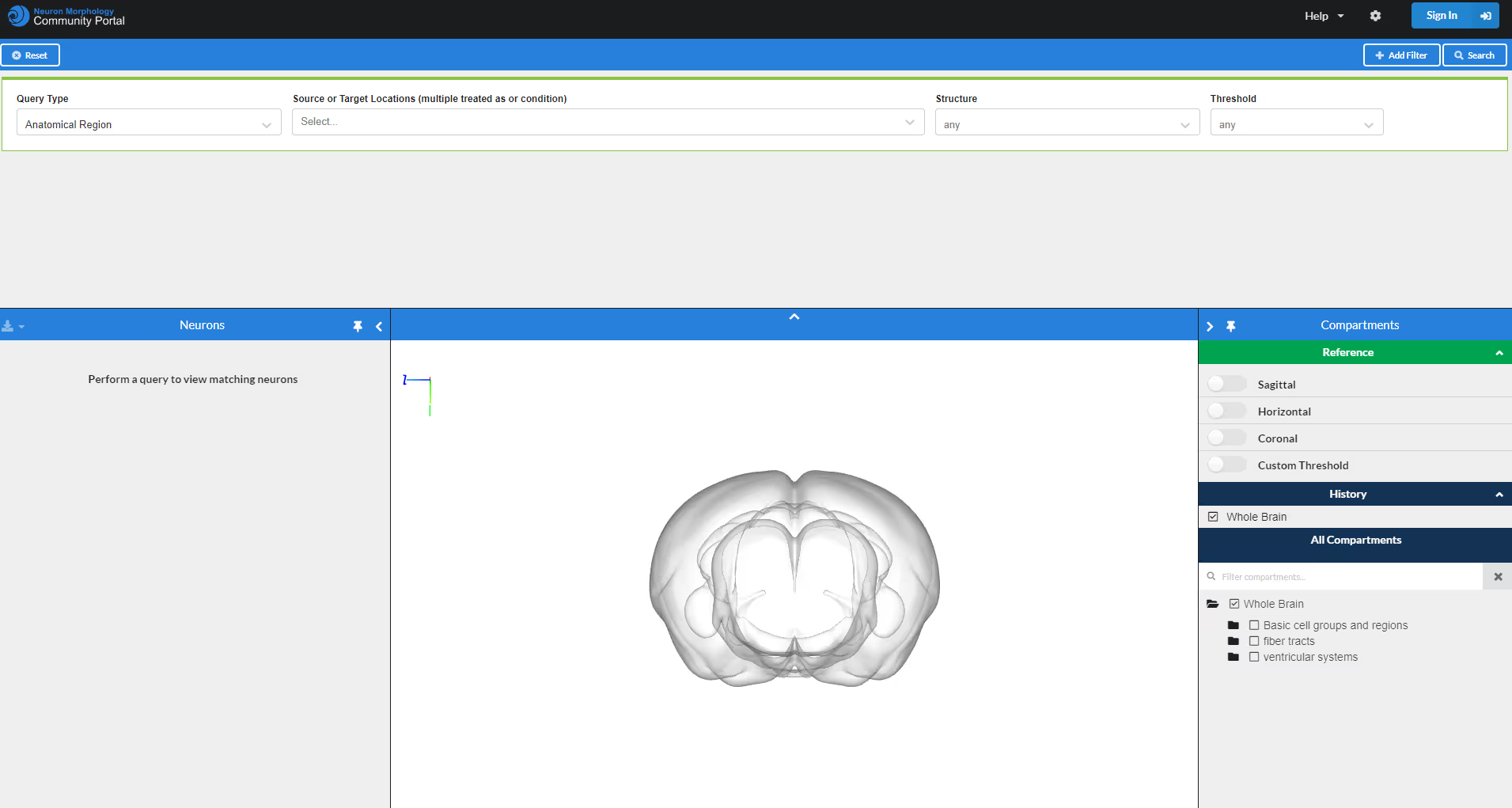
The Neuron Morphology Community Portal is is hub to search for whole-brain reconstructions of neuron morphologies in the adult mouse brain. It can host any data that has been aligned to the Allen Mouse Brain Common Coordinate Framework. Log into contribute new neuron skeletons or sign up to proofread new neurons with HortaCloud.
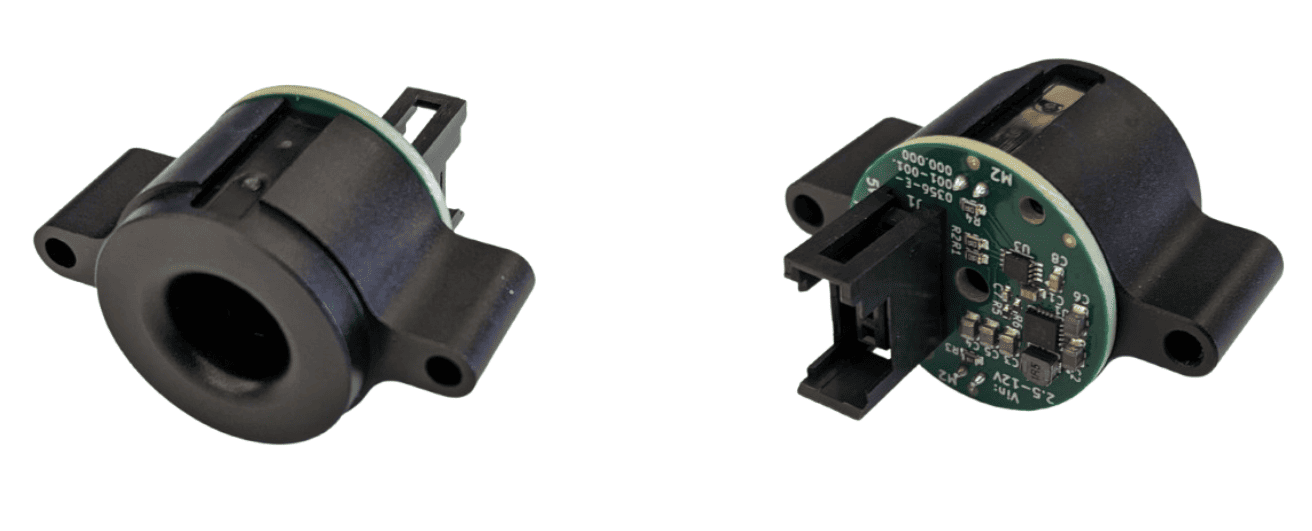

Harp-compliant poke port for freely moving behavior.
Focusing on multicamera video streaming for microscopy.
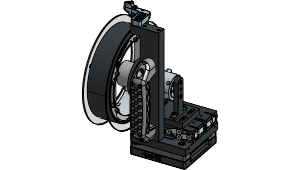

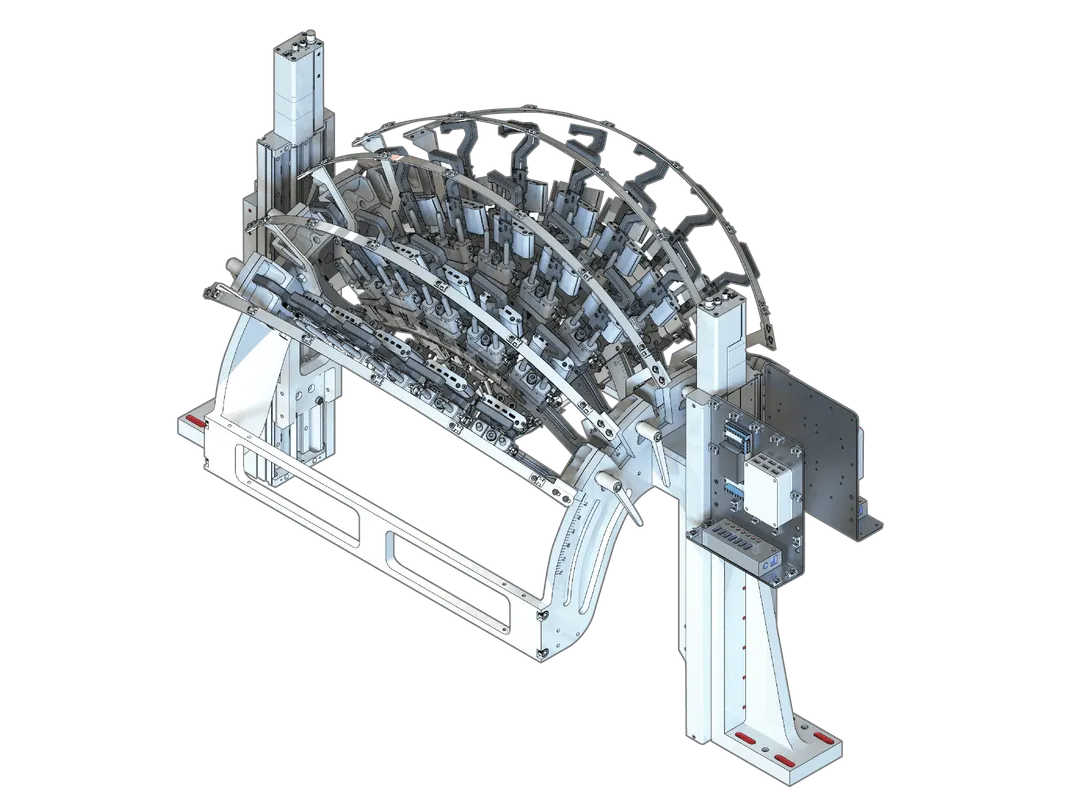
New treadmill-style running wheel enables natural running posture, closed-loop variable effort running, headframe integration for a range of neurophysiology experiments, and measuring torque and speed of wheel.
.png)
Headframe templates and designs for the Allen Institute headframe clamp system for intersession and interinstrument spatial registration.

Compact, cost-effective, modular stimulus tool stage suitable for mouse behavior tasks.

Single and dual hemisphere implant templates with supporting installation tools.
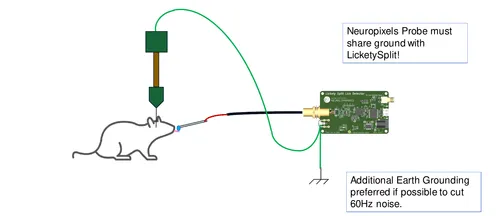
Signal conditioning board for an implanted thermistor.
An ephys-compliant capacitive lick detector.
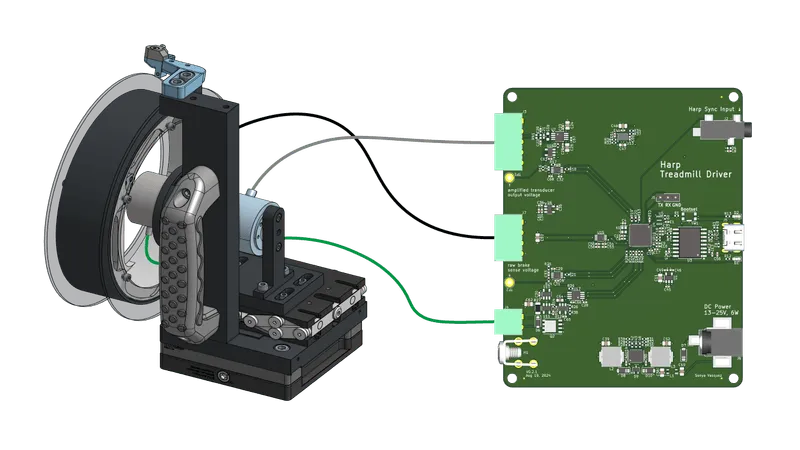
A Harp-compatible device for controlling an instrumented mouse treadmill.
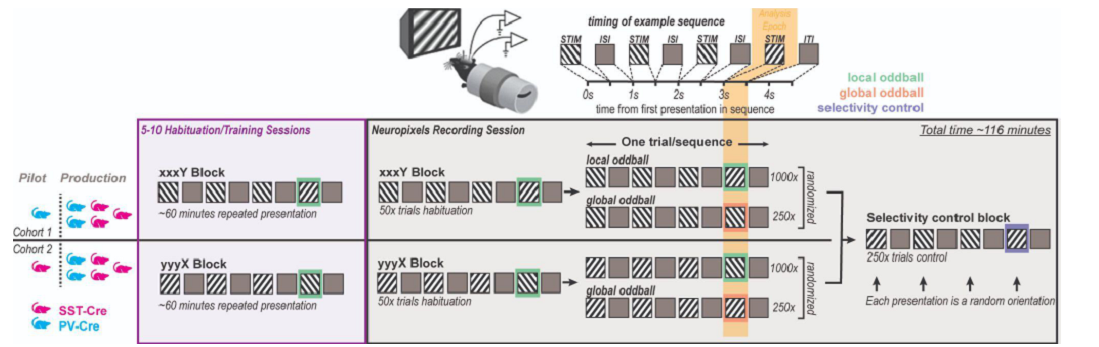
This dataset was collected for the Global Local Oddball project, as part of the OpenScope project.
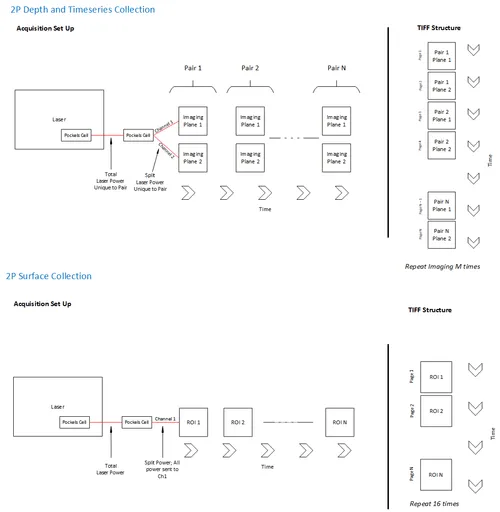
The multiplane pipeline processes planar optical physiology data acquired in parallel to extract events from the ROIs in each plane.
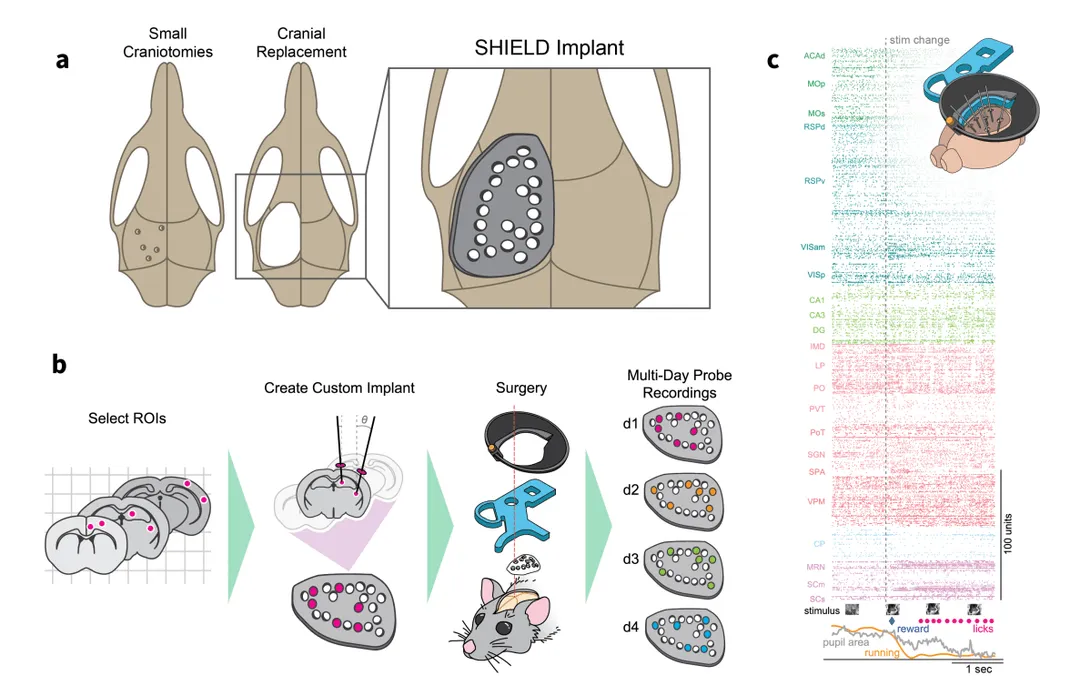
We describe a novel 3D-printed cranial-replacement implant (SHIELD) enabling electrophysiological recordings from distributed areas of the mouse brain, which allows us to measure spiking dynamics across many interacting brain regions.

This high magnification lens is a specialist for light sheet microscopy with a very large field of view. With its magnification of 5x and its image circle of 82mm it permits the analysis of larger samples in a single image, obviating the necessity for additional segmentation.

The ExA-SPIM method represents a major breakthrough in microscopy. Schneider-Kreuznach's DIAMOND lenses play a crucial role. Their large image circle and high magnification allow the analysis of larger samples in a single image without additional segmentation.

A library that defines AIND data schema and validates JSON.

This protocol describes the delipidation of a mouse brain specimen using a modified iDISCO protocol.
This service can be used to upload data stored in a VAST drive. It uses FastAPI to upload a job submission csv file that will be used to trigger a data transfer job in an on-prem HPC. Based on the information provided in the file, the data upload process fetches the appropriate metadata and starts the upload process.
This protocol describes the pre-operative setup and post-operative take-down procedures utilized for rodent stereotaxic neurosurgical procedures.

This protocol describes the steps necessary to apply this silicone layer in advance of a cranial windowing procedure.
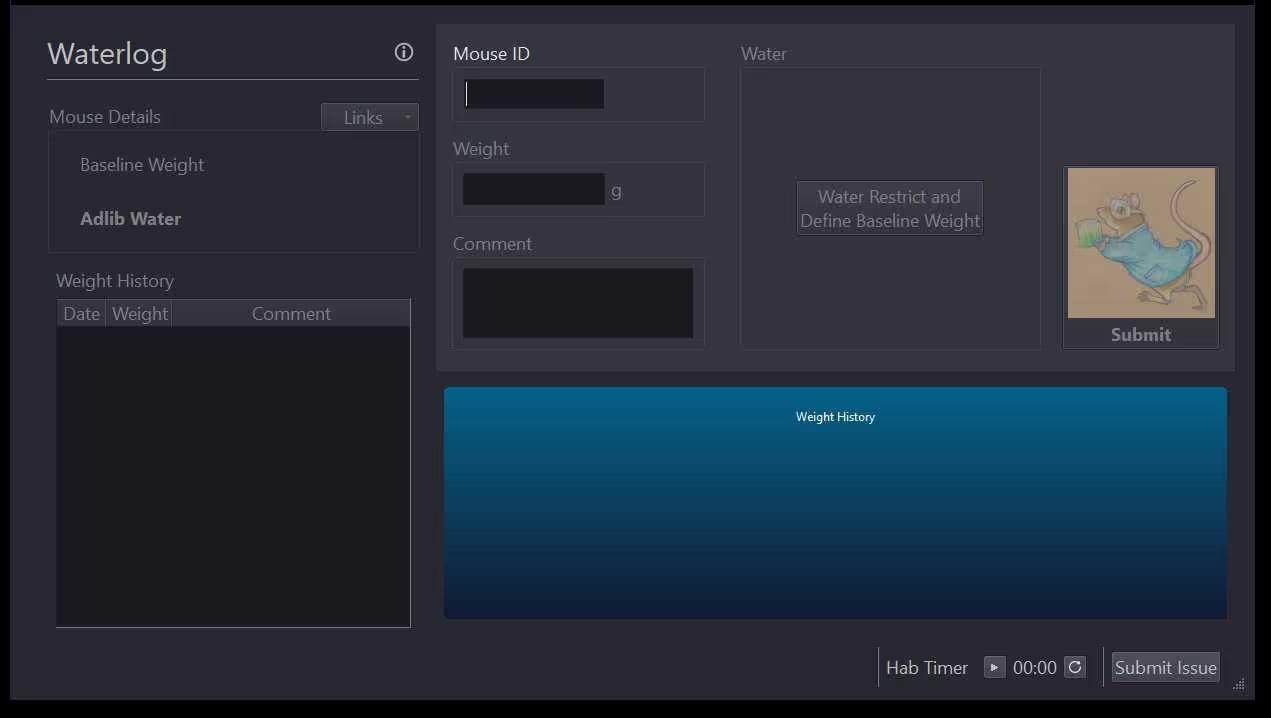
This protocol describes a semi-customized workflow to perform water restriction with mice designated for future experiments.
.avif)
This protocol describes a customized mouse habituation protocol for mice within experiments involving head fixation onto disks.
(1).webp)
This protocol describes the procedure for removing the SORTA-clear plug used in acute in vivo electrophysiology experiments in whole-hemisphere craniotomy (WHC) preparation mice with Neuropixels probes.
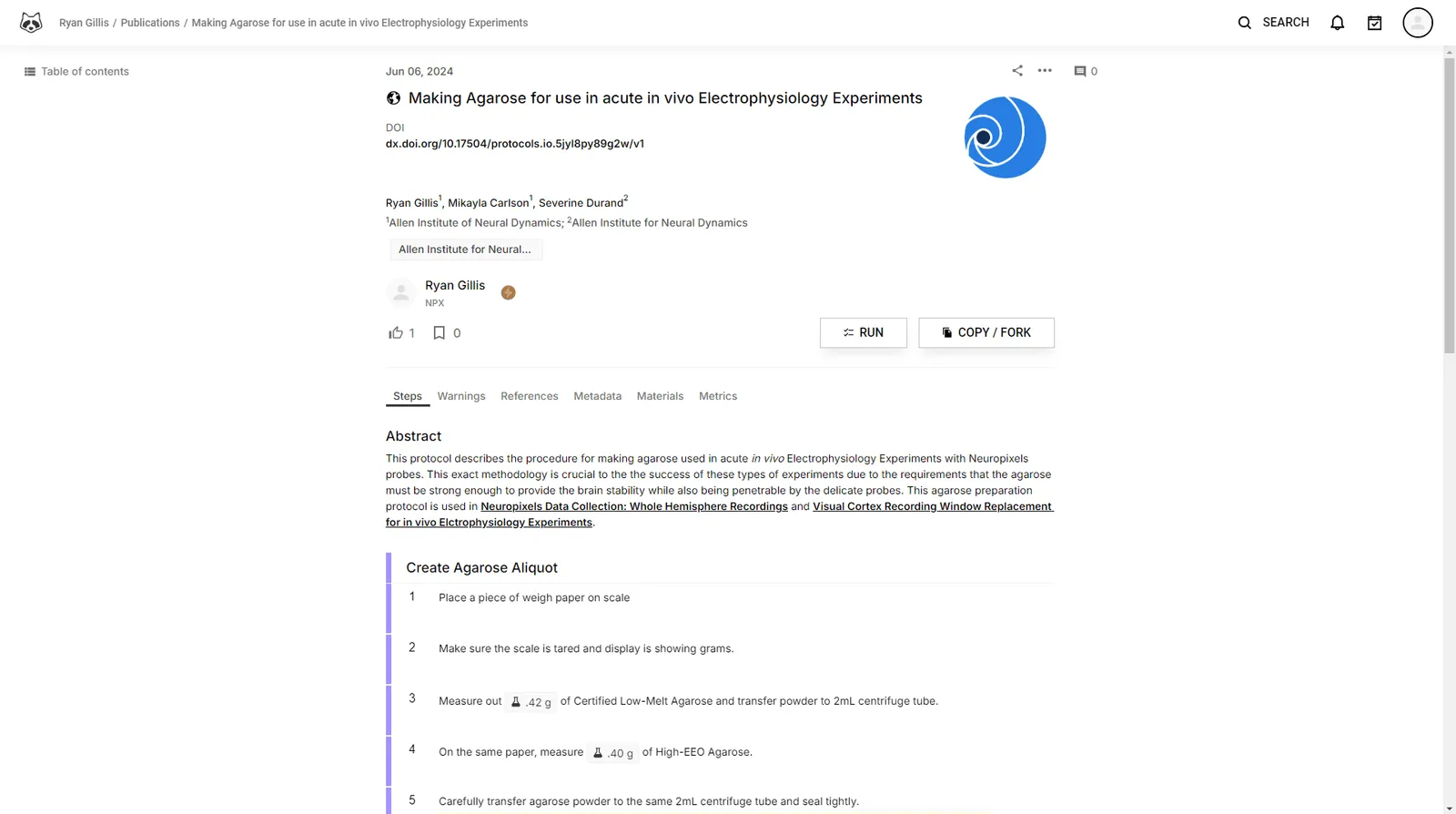
This protocol describes the procedure for making agarose used in acute in vivo Electrophysiology Experiments with Neuropixels probes.
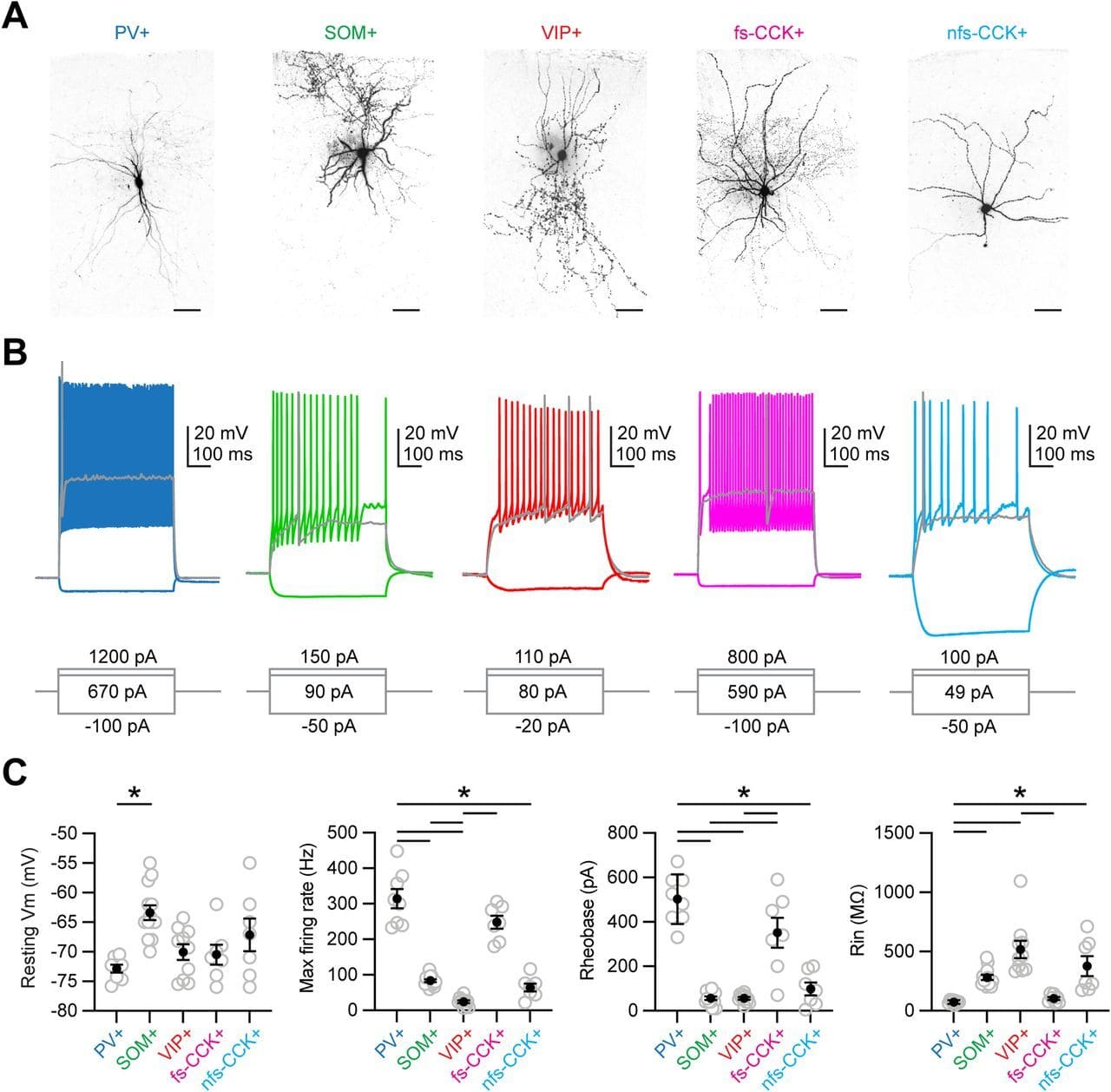
Here, we use anatomy, slice physiology, and optogenetics to examine how mediodorsal (MD) thalamus evokes feed-forward inhibition in Layer 3 of the prelimbic subregion of the mouse prefrontal cortex (PFC).
%20(1).jpg)
We study the biological mechanisms that enable animals to learn the structure of their world through free exploration. To do this, we collect long-term, uninterrupted records of the natural behavior, brain activity, and network connectivity of mice while they repeatedly interact with odors in their environment.
.png)
We are developing and using genetic, electrophysiological, optical, and behavioral approaches to investigate how the brain adaptively controls behavior. The team focuses on understanding the descending circuits that control the execution of actions and how they change when actions are reinforced and refined.
We are using large-scale electrophysiology to study how distributed brain regions coordinate their spiking activity to guide behavior in changing environments.