Mesoscale Anatomy
.png)
.png)
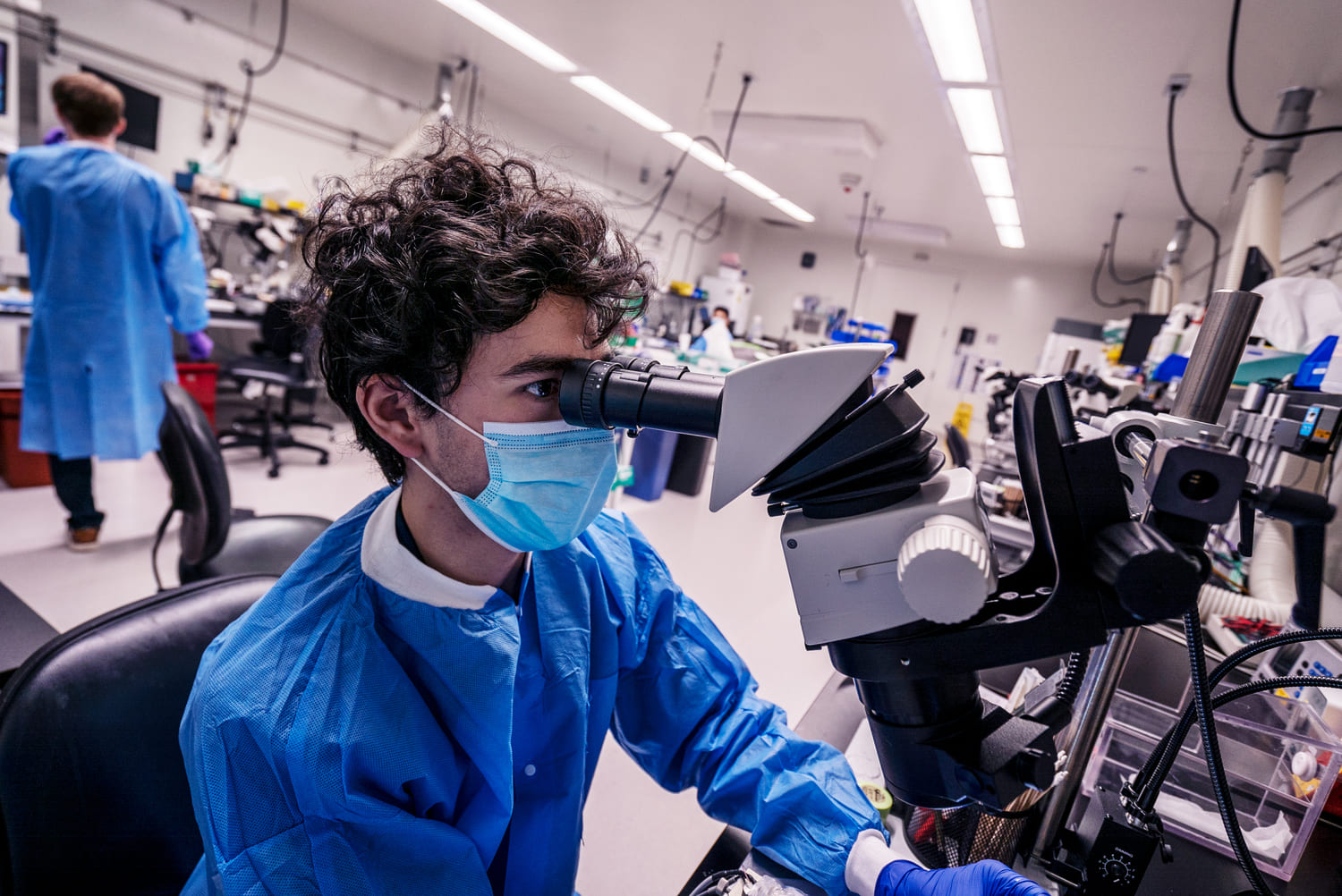
Neural dynamics are generated and constrained by connections between neurons in both local and brain-wide circuits. Understanding the anatomical channels by which information flows through the brain is thus critical to understanding neural activity and computation. Furthermore, placing neurophysiological recordings or newly developed molecular tools in the context of the brain’s anatomical organization is key for interpreting brain-wide activity. The Mesoscale Anatomy platform is a tissue and data processing pipeline for brain-wide histology, enabling routine, reproducible, and quantitative studies.
The Mesoscale Anatomy platform relies upon tissue clearing technology to render whole-brains optically transparent, often while preserving native fluorescence (e.g. from transgenic or virally expressed fluorophores).This enables volumetric imaging of whole brains at cellular resolution, without slicing or otherwise damaging tissue. Imaged brains create terabytes of data, and a computational pipeline transforms “data” into “insight” via automated end-to-end workflows for image-processing, registration, and cell segmentation. This platform enables standardized whole-brain histology for use cases including (but not limited to) characterizing molecular tools, localizing neurophysiology probes, and viral tracing for quantitative connectivity studies.
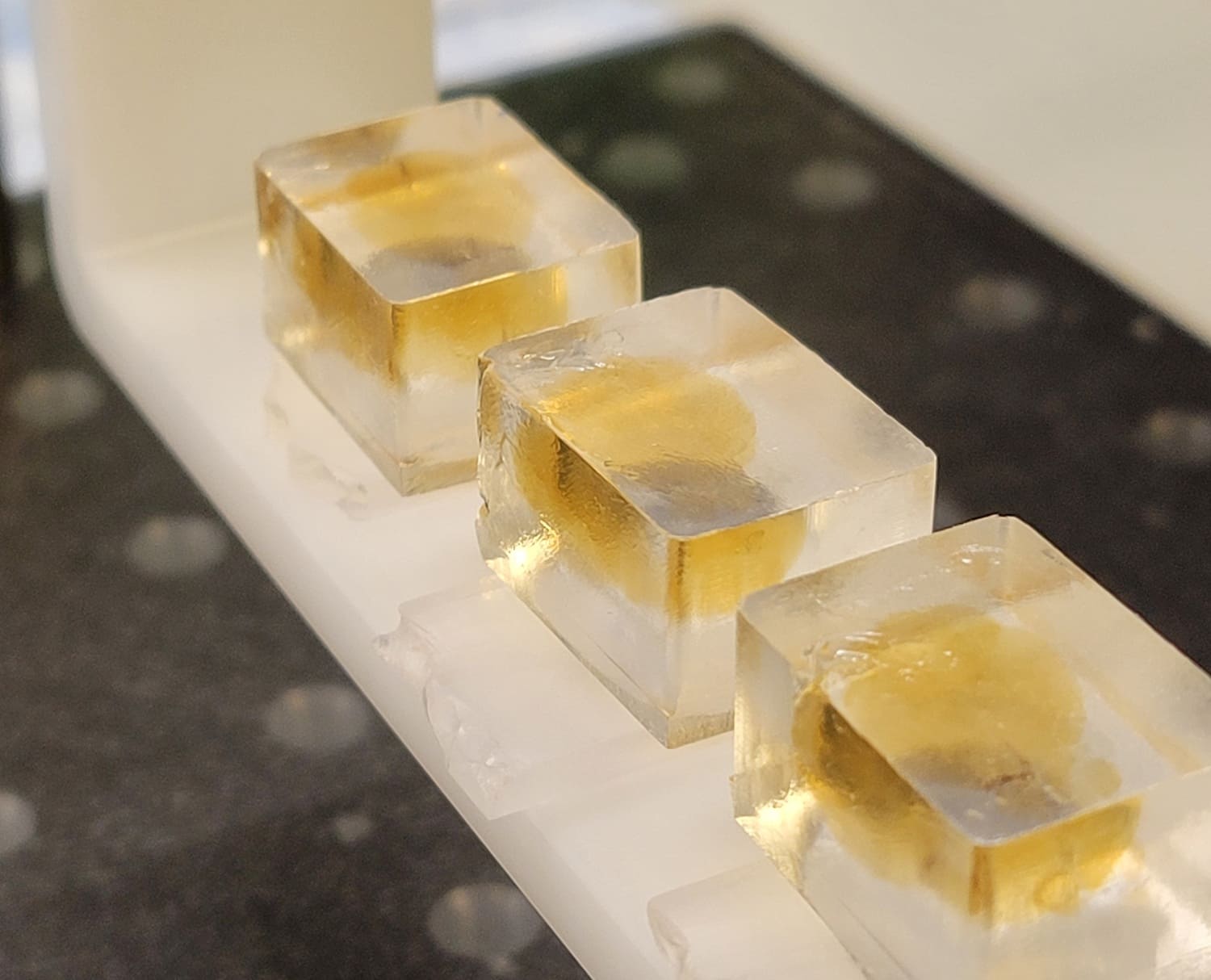
Volumetric imaging of intact tissue requires tissue clearing to render brains optically transparent. These techniques involve delipidating and index matching biological samples to allow light to penetrate the brain with minimal diffraction or scattering. The Mesoscale Anatomy platform utilizes a variety of tissue clearing protocols tailored to the needs of a given experiment. Most experiments rely on aqueous based clearing techniques, preserving native fluorescence and reducing the need for subsequent signal amplification via antibody labeling.
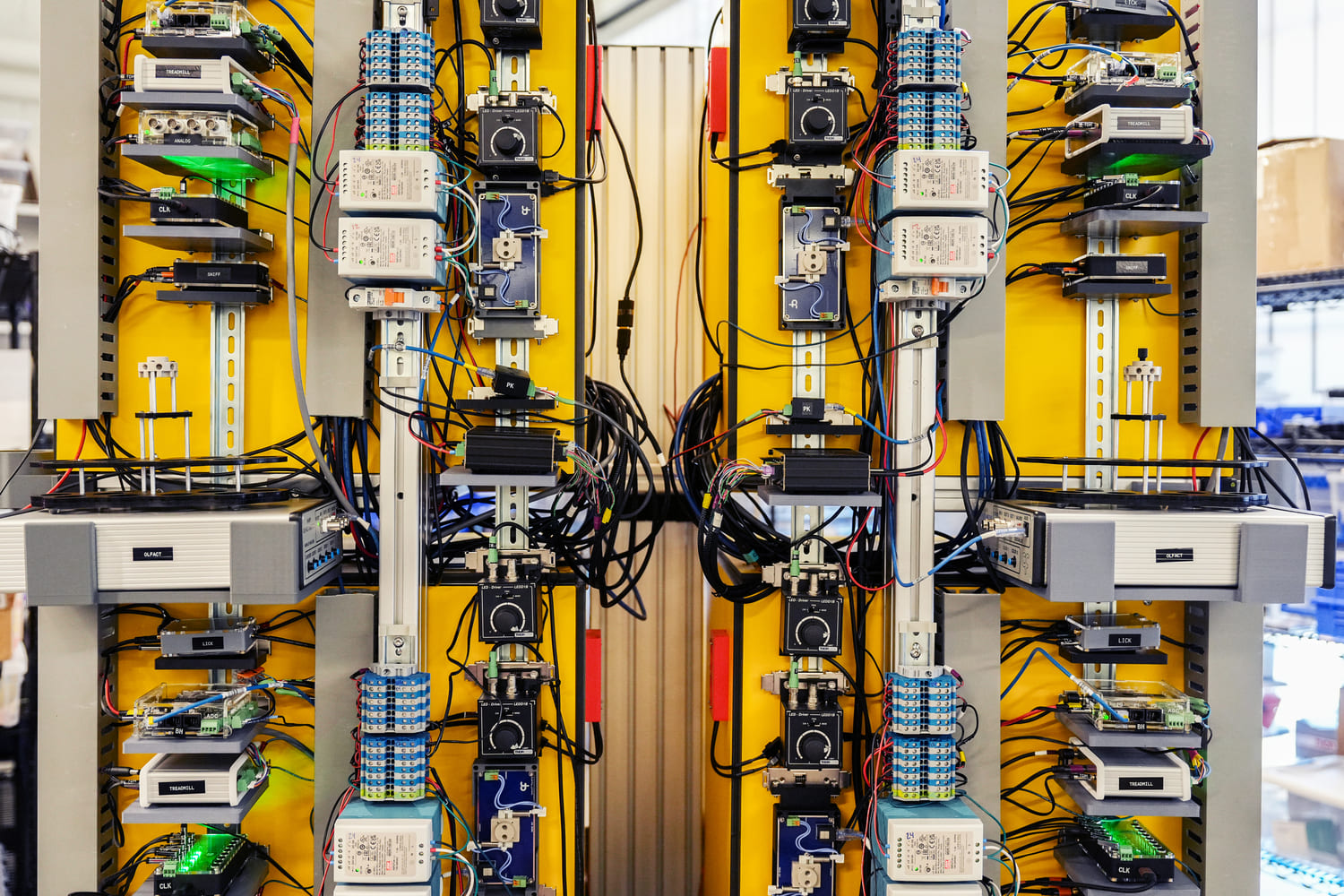
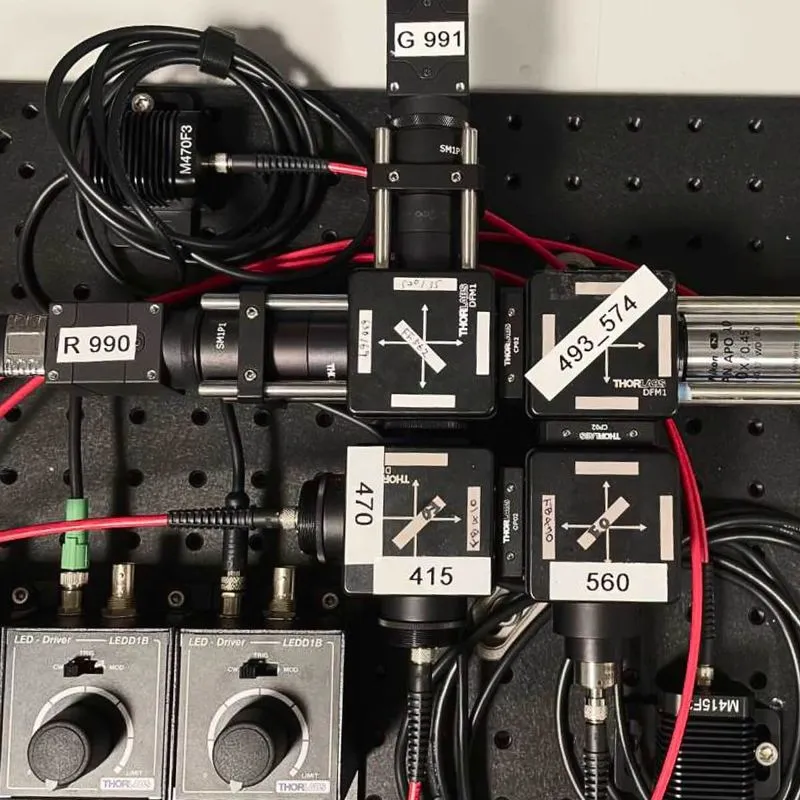
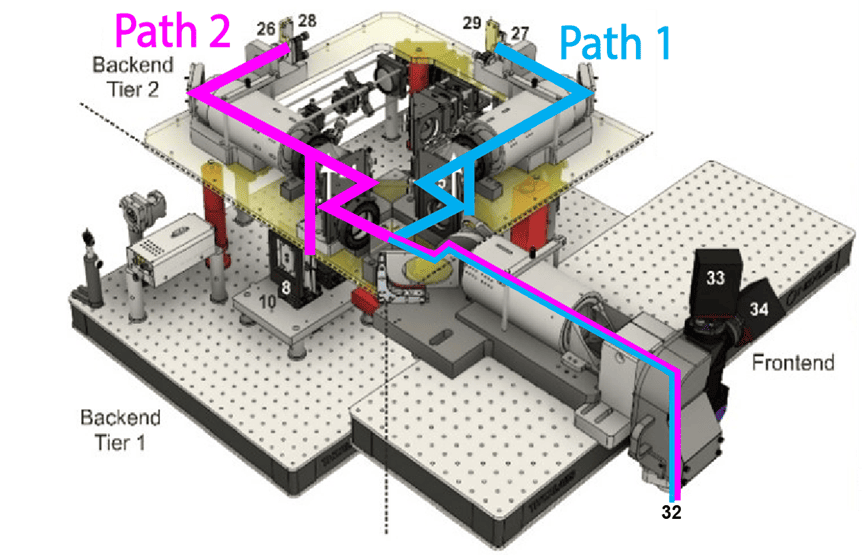
Samples are then imaged nondestructively on a selective plane illumination microscope (SPIM), which allows rapid acquisition of volumetric data. Mouse brains are imaged at 1.8 x 1.8 x 2.0 micron resolution, at a rate of approximately 2 hours per spectral channel. The Mesoscale Anatomy team processes dozens of brains per week, and regularly outputs over a thousand brains a year.
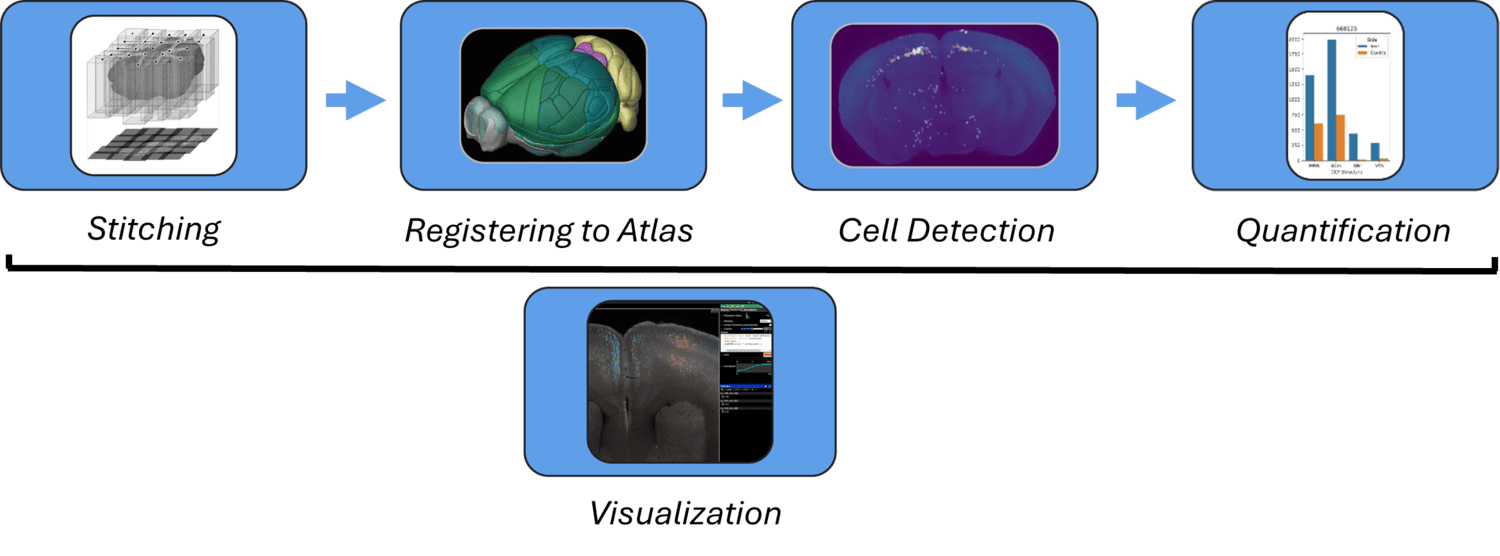
The Mesoscale Anatomy platform supports an automated end-to-end image processing pipeline to transform large datasets into biological insights. Modular compute capsules are used to support different experimental use cases. After imaging, data undergoes pre-processing such as flatfield correction and destriping. Images are then stitched and fused to generate 3D volumes. Brains are subsequently registered to CCF, which aligns distinct experiments into a shared coordinate space for unified analyses. Optional post-processing modules also support automated cell detection via machine vision. The data products from each step of this pipeline are programmatically accessible, as well as visualizable via web-browser based Neuroglancer views (e.g. https://tinyurl.com/72fh2m72). Computing workflows are optimized for use within the Code Ocean computing platform, but they are also deployable on local clusters or individual machines.
Integrating analyses between experiments requires aligning samples to a common reference space. The Allen Mouse Brain Common Coordinate Framework (CCF)1 is a 3D atlas that allows for co-registration of experimental samples and provides anatomical annotation of brain structures. Cross-modal alignment of whole-brain SPIM datasets to the CCF (originally generated via serial two-photon tomography) is achieved using an intermediate SPIM template created by averaging many cleared brains. Automated CCF registration enables robust, reproducible science and anatomically informed analyses of large experimental cohorts.
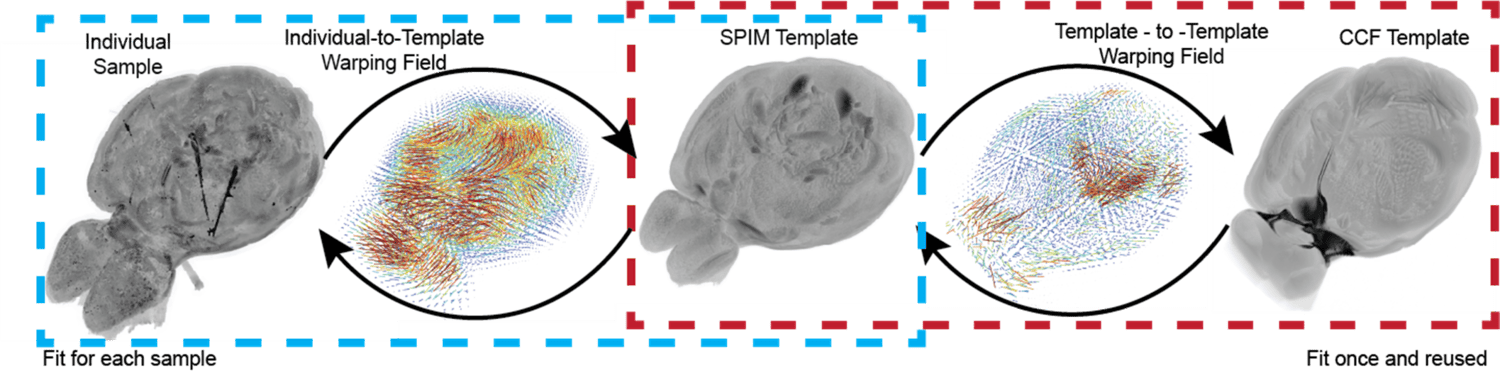
1Wang, Q., Ding, S., Li, Y., et al. The Allen Mouse Brain Common Coordinate Framework: A 3D Reference Atlas. Cell 181, 936-953.e20 (2020), https://doi.org/10.1016/j.cell.2020.04.007.
Machine vision workflows are crucial for sifting through terabyte-scale datasets for biologically meaningful signals in an efficient and reproducible manner. Automated cell detection identifies the number and location of cells across the brain.2 These cell detection tools enable quantitative analysis for a variety of experimental use cases, including molecular tool characterization, connectivity studies utilizing viral tracing, and activity mapping via immediate early gene expression.
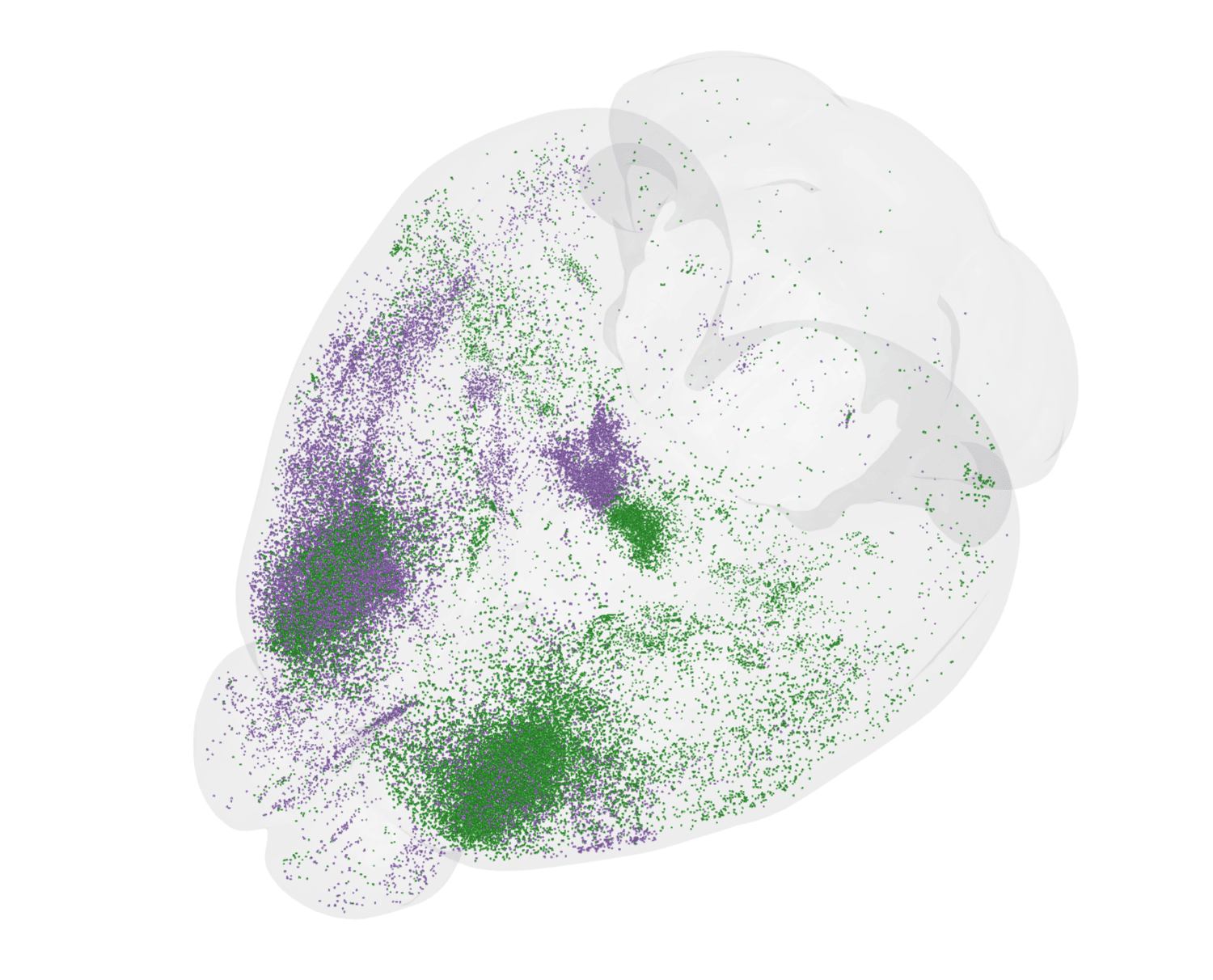
2Tyson, A. L. et al. A deep learning algorithm for 3D cell detection in whole mouse brain image datasets. PLOS Computational Biology 17, e1009074 (2021), https://doi.org/10.1371/journal.pcbi.1009074.
This platform builds on turn-key lightsheet microscopes to image mouse brains at high resolution and throughput. Analysis of the resulting volumetric images, including registration to standard brain coordinates and segmentation and counting of individual neurons, is fully automated.
This platform uses a two-photon microscope SLAP2 that records patterns of synaptic input and output in individual neurons at hundreds to thousands of frames per second in mice performing complex behaviors, using a flexible scan system that combines a digital micromirror device (DMD) with a high-speed scanner.
The Scientific Instrumentation and Process Engineering (SIPE) team is a shared engineering resource within the Allen Institute, focused on enabling and scaling cutting-edge bioscience through integrated hardware and software systems.
This team performs a variety of surgical procedures, including stereotaxic injections and implanting chronic cranial windows and Neuropixels probes.
This platform implements pioneering technology for highly reproducible, targeted, brain-wide, cell-type-specific electrophysiology to record neural activity from defined neuron types across the brain. Analysis and quality control of the electrophysiology data are fully automated.
This platform combines innovative histology, ExA-SPIM microscopy, image handling, and machine learning to map the morphology and molecular identity of individual neurons across the whole brain at high throughput.
This platform enables optical measurement of neural activity and neurotransmitter release in populations of neurons to study neural circuit dynamics in behaving animals.
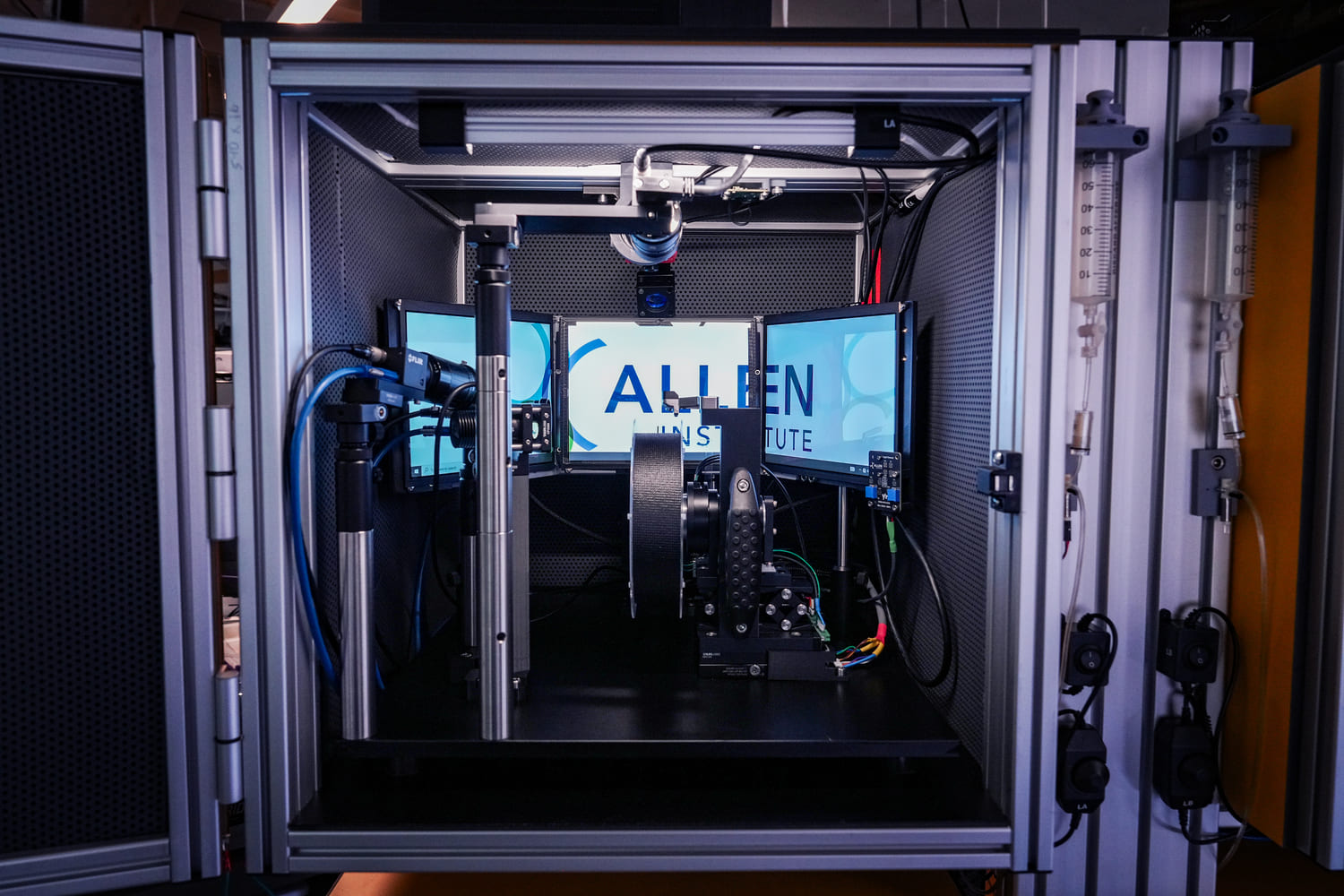
The Behavior platform uses advanced technology to implement a standardized, modular, multi-task virtual reality gymnasium for mice, with the goal to study brain function across different behaviors at scale.