SLAP2


The living brain is opaque: it absorbs and scatters light. Most imaging methods rapidly lose resolution and sensitivity deeper than tens of microns below the brain surface. To image tiny features deeper inside the brain, neuroscientists use two-photon microscopy, in which pulsed infrared laser light excites fluorescence one voxel at a time.
This serial excitation process is fundamentally slow. Each voxel takes at least a few nanoseconds to measure, a limit imposed by the time it takes an excited fluorophore to emit light. The volume containing just a single neuron’s dendrites can include on the order of a billion resolvable voxels.
Measurement speed is critically important for studying how neural circuits process information. Neural activity evolves on millisecond timescales, and detecting coincidences in inputs from two different synapses onto a neuron can require sampling at >100 Hz.
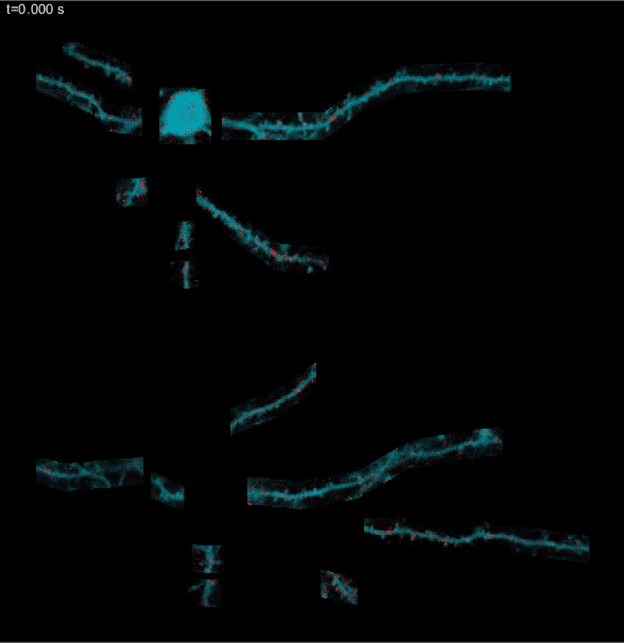
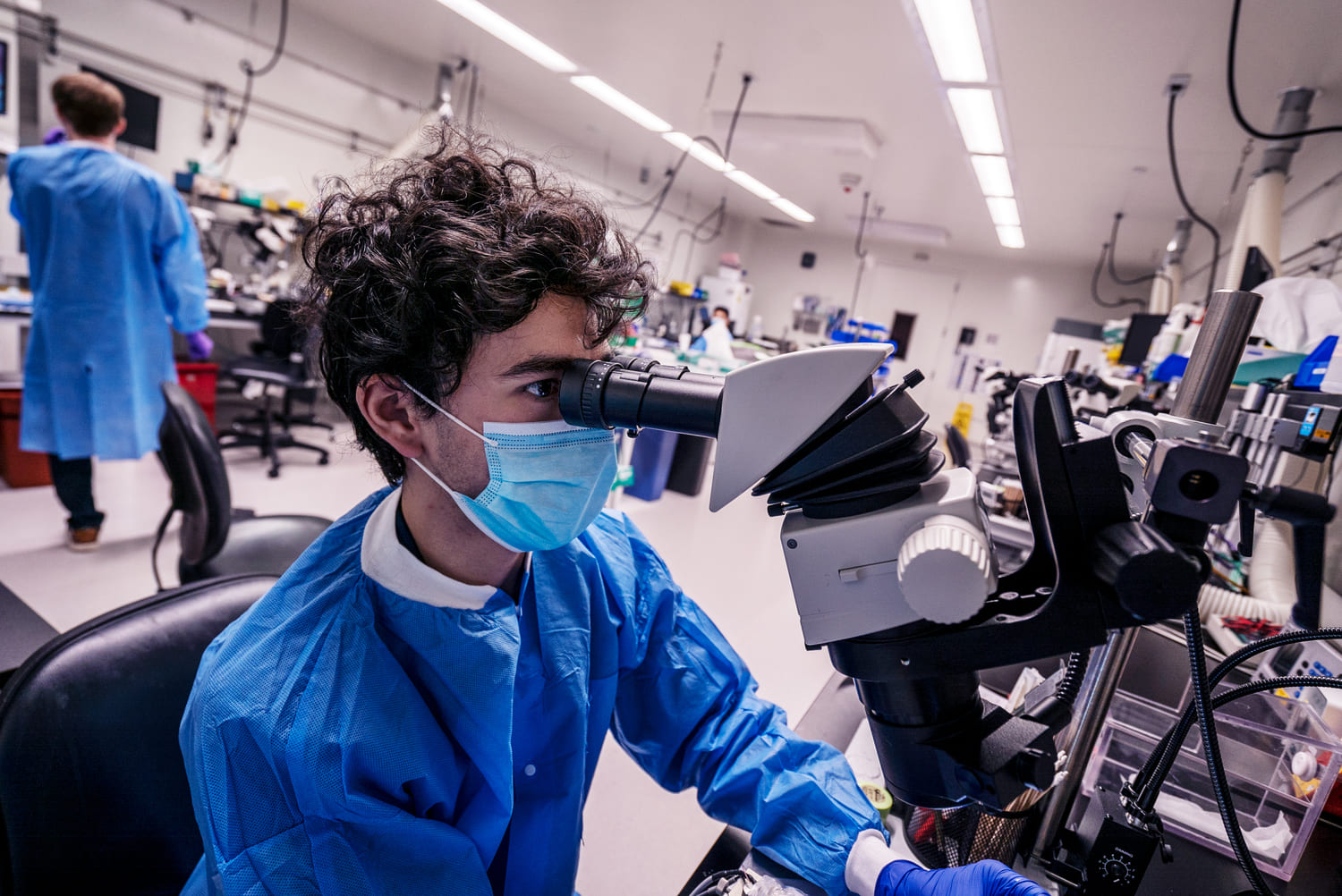
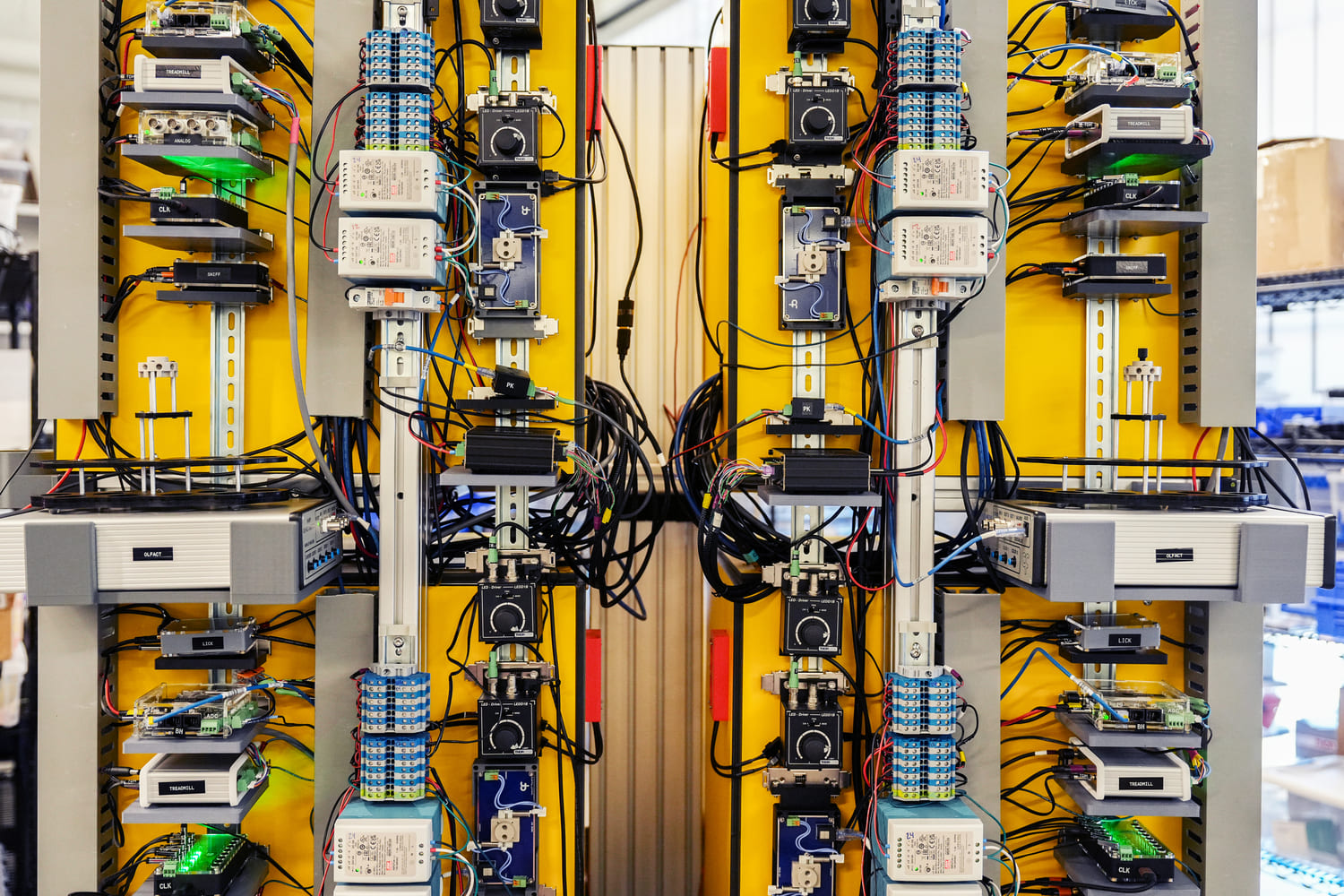
SLAP2 is a two-photon microscope based on a new scanning system designed to address these challenges. It records patterns of synaptic input (e.g., glutamate) and output (e.g., membrane voltage) in individual neurons at hundreds to thousands of frames per second in mice performing complex behaviors.
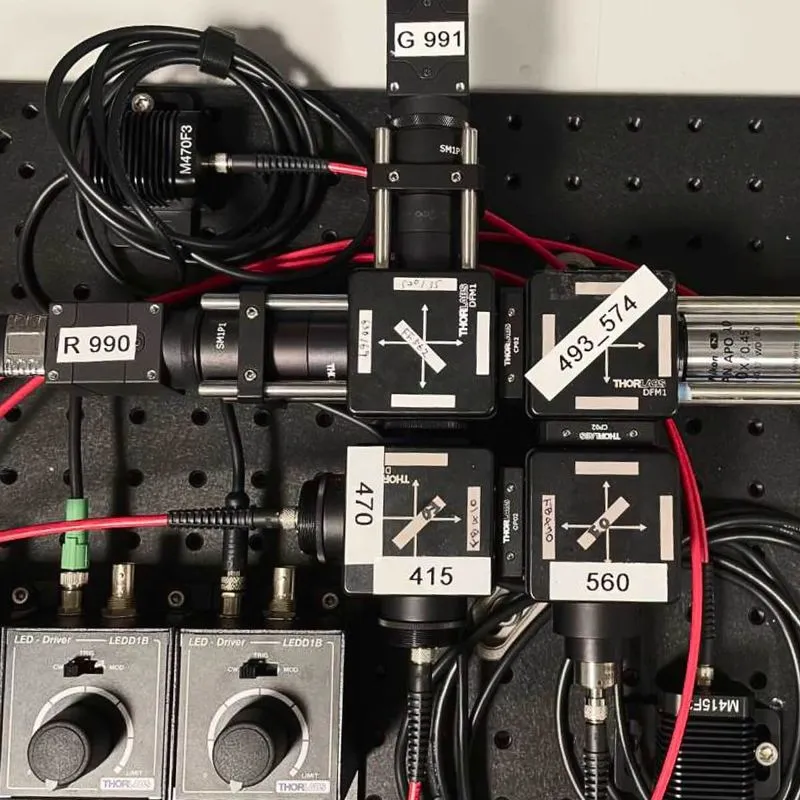
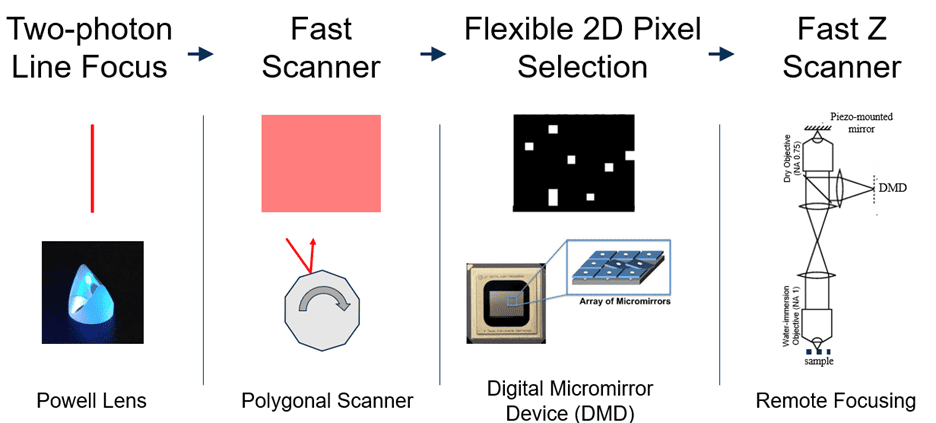
SLAP2 uses a flexible scan system that combines a digital micromirror device (DMD) with a high-speed scanner. This lets it target only selected pixels for imaging, with frame time proportional to the number of pixels selected—an approach often called random-access imaging. Previous random-access microscopes required access times of tens of microseconds to jump between imaging targets (thousands of times longer than the time to acquire a single pixel). In contrast, SLAP2 has no per-target access-time cost.
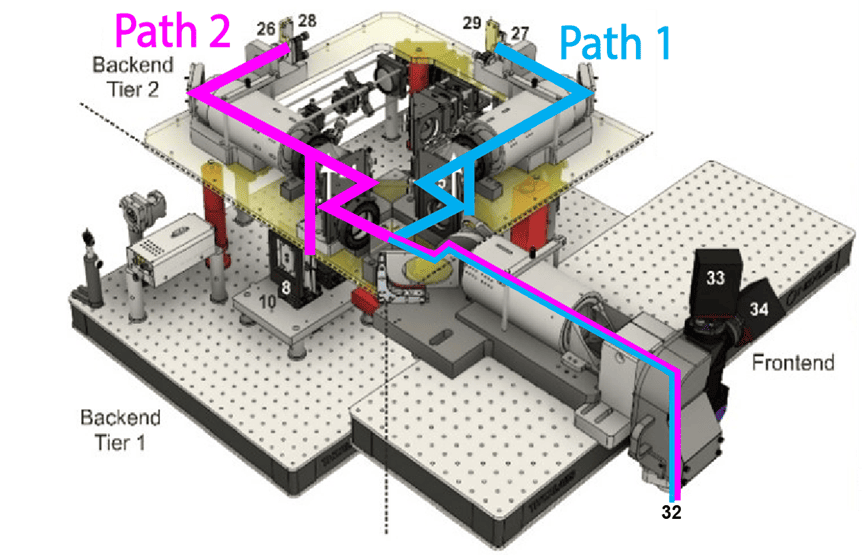
SLAP2 uses two independent DMD-based imaging paths that are independently steered, enabling recordings of hundreds of synapses from multiple neuron regions at once.
SLAP2 microscope kits are available from MBF Bioscience.
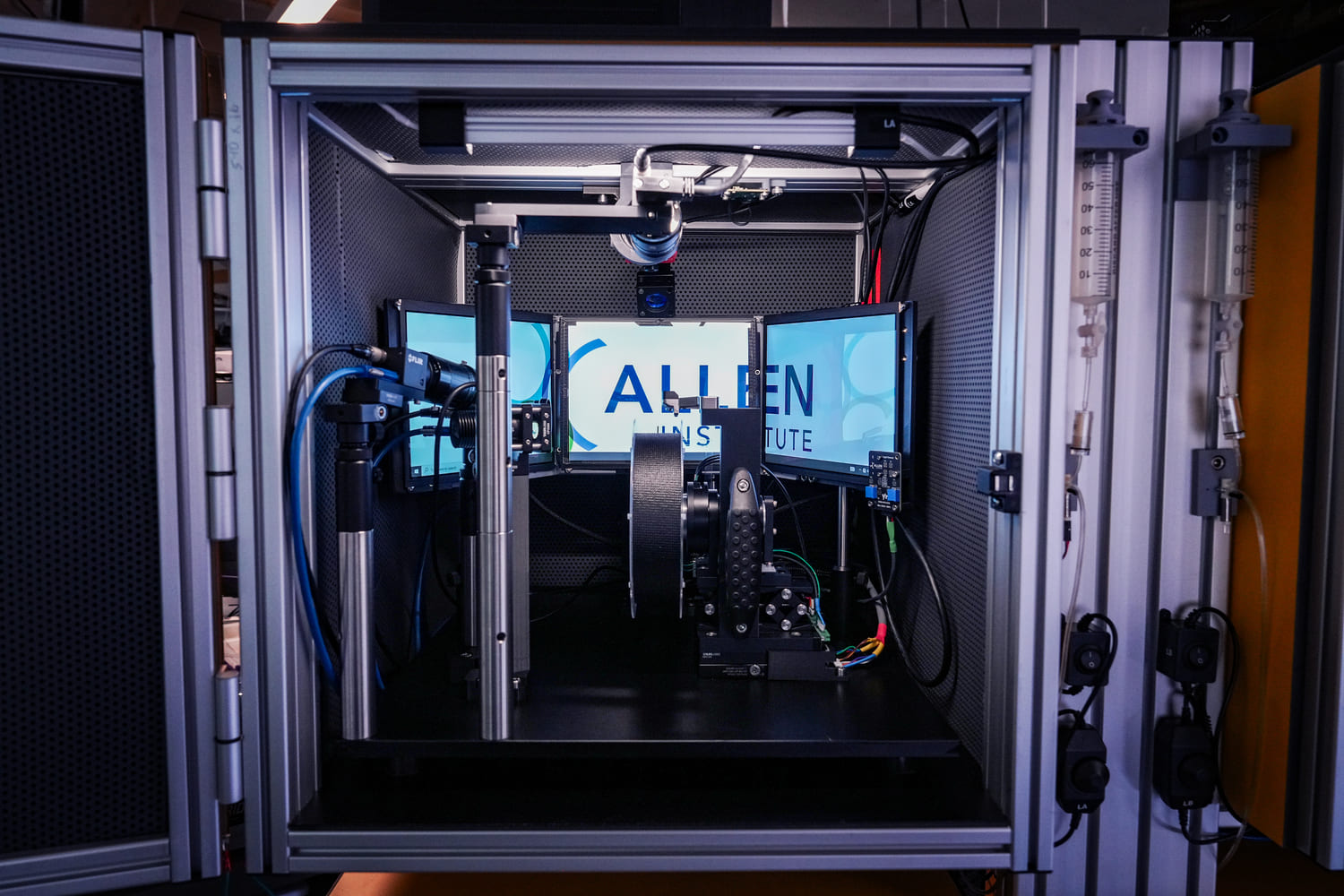
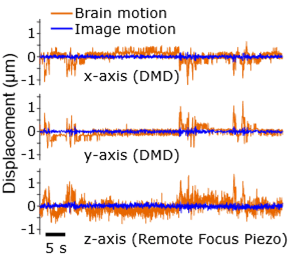
During imaging, SLAP2 tracks brain movement from the incoming image stream and corrects it in real time, with latencies under 100 microseconds. This compensates for most motion artifacts, even in head-fixed mice running on a treadmill.
To visualize inputs to neurons, we develop fluorescent protein neurotransmitter indicators. Neurons communicate at synapses by releasing brief (~1 ms) bursts of neurotransmitter molecules, such as glutamate, GABA, or dopamine. To make these molecules visible, we engineer genetically encoded indicators that bind neurotransmitters and fluoresce. These indicators are expressed on a neuron’s surface to report its synaptic inputs.
In collaboration with other protein-engineering groups, we are developing next-generation indicators for a variety of neurotransmitters and neuromodulators. Our glutamate indicators, iGluSnFR3 and iGluSnFR4, are available from Addgene.
Spontaneous glutamate release in cultured neurons expressing iGluSnFR3. These neurons have been silenced with TTX. The red flashes are miniature release events caused by asynchronous, spontaneous release of individual synaptic vesicles. Detecting these fundamental quanta of neural communication highlights the high sensitivity of this indicator.
SLAP2 has multiple imaging modes that trade off speed,scale, and resolution. It can record hundreds of synapses at >200 Hz inmulti-ROI raster mode, and >1,000 synapses at >50 Hz in band-scanningmode. Integration mode enables multi-kHz random-access voltage imaging frommany targets at once.
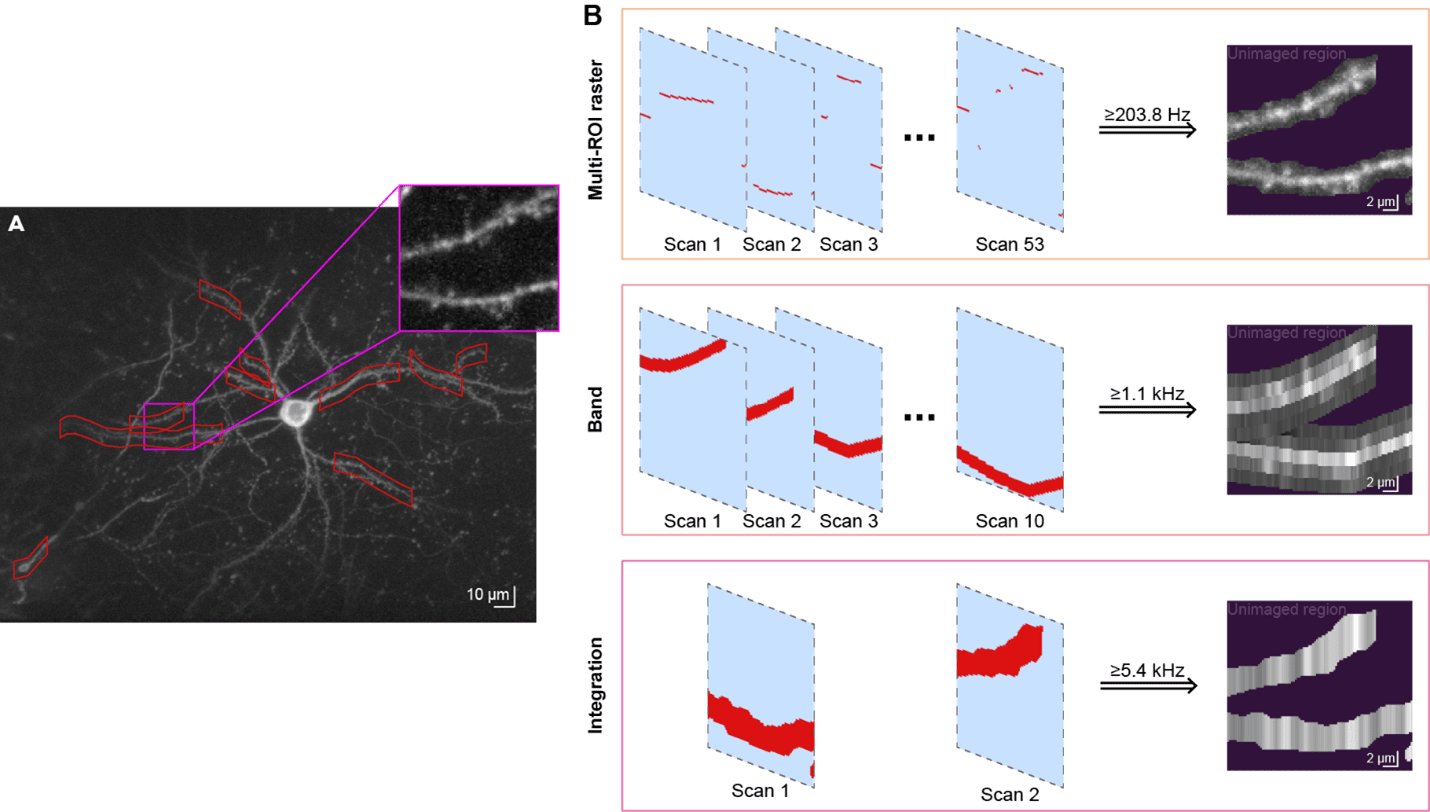
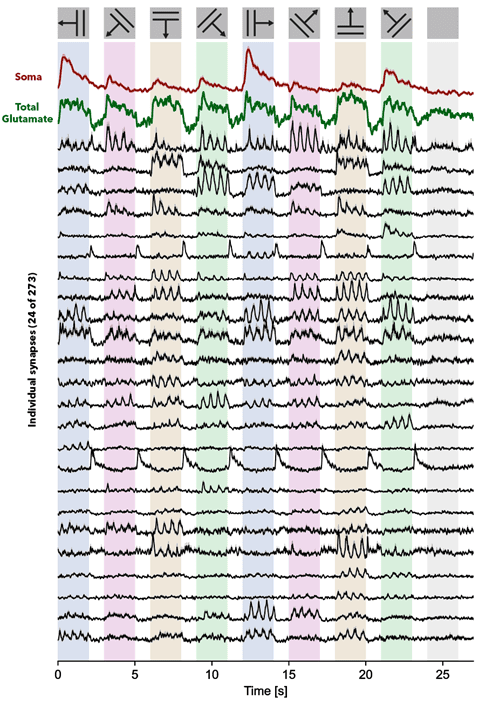
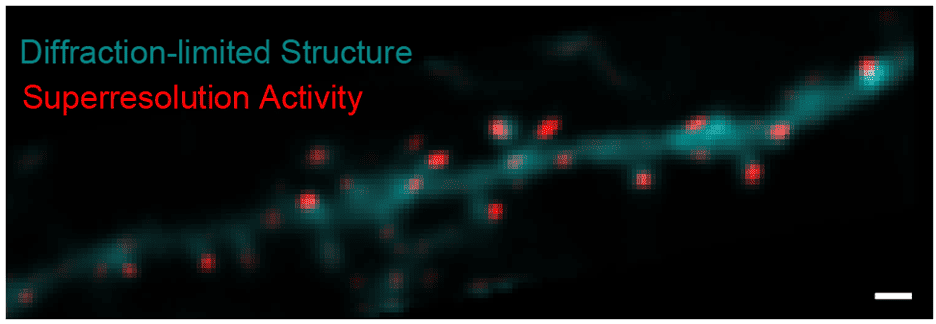
Incoming image data from photodetectors is processed to extract biologically relevant signals, such as synaptic activity. High-speed imaging of fast indicators allows us to adapt methods from localization microscopy to generate super-resolution maps of activity that report synapse locations. Using the inferred synapse locations, we apply matrix-factorization approaches to decompose the recordings into the activity of each synapse overtime. We evaluate data-processing algorithms using multiple ground-truth strategies, including post hoc tissue expansion and immunohistochemical labeling of synapses.

Additional funding from:
This platform builds on turn-key lightsheet microscopes to image mouse brains at high resolution and throughput. Analysis of the resulting volumetric images, including registration to standard brain coordinates and segmentation and counting of individual neurons, is fully automated.
This platform uses a two-photon microscope SLAP2 that records patterns of synaptic input and output in individual neurons at hundreds to thousands of frames per second in mice performing complex behaviors, using a flexible scan system that combines a digital micromirror device (DMD) with a high-speed scanner.
The Scientific Instrumentation and Process Engineering (SIPE) team is a shared engineering resource within the Allen Institute, focused on enabling and scaling cutting-edge bioscience through integrated hardware and software systems.
This team performs a variety of surgical procedures, including stereotaxic injections and implanting chronic cranial windows and Neuropixels probes.
This platform implements pioneering technology for highly reproducible, targeted, brain-wide, cell-type-specific electrophysiology to record neural activity from defined neuron types across the brain. Analysis and quality control of the electrophysiology data are fully automated.
This platform combines innovative histology, ExA-SPIM microscopy, image handling, and machine learning to map the morphology and molecular identity of individual neurons across the whole brain at high throughput.
This platform enables optical measurement of neural activity and neurotransmitter release in populations of neurons to study neural circuit dynamics in behaving animals.
The Behavior platform uses advanced technology to implement a standardized, modular, multi-task virtual reality gymnasium for mice, with the goal to study brain function across different behaviors at scale.