Tools, Publications, & Data


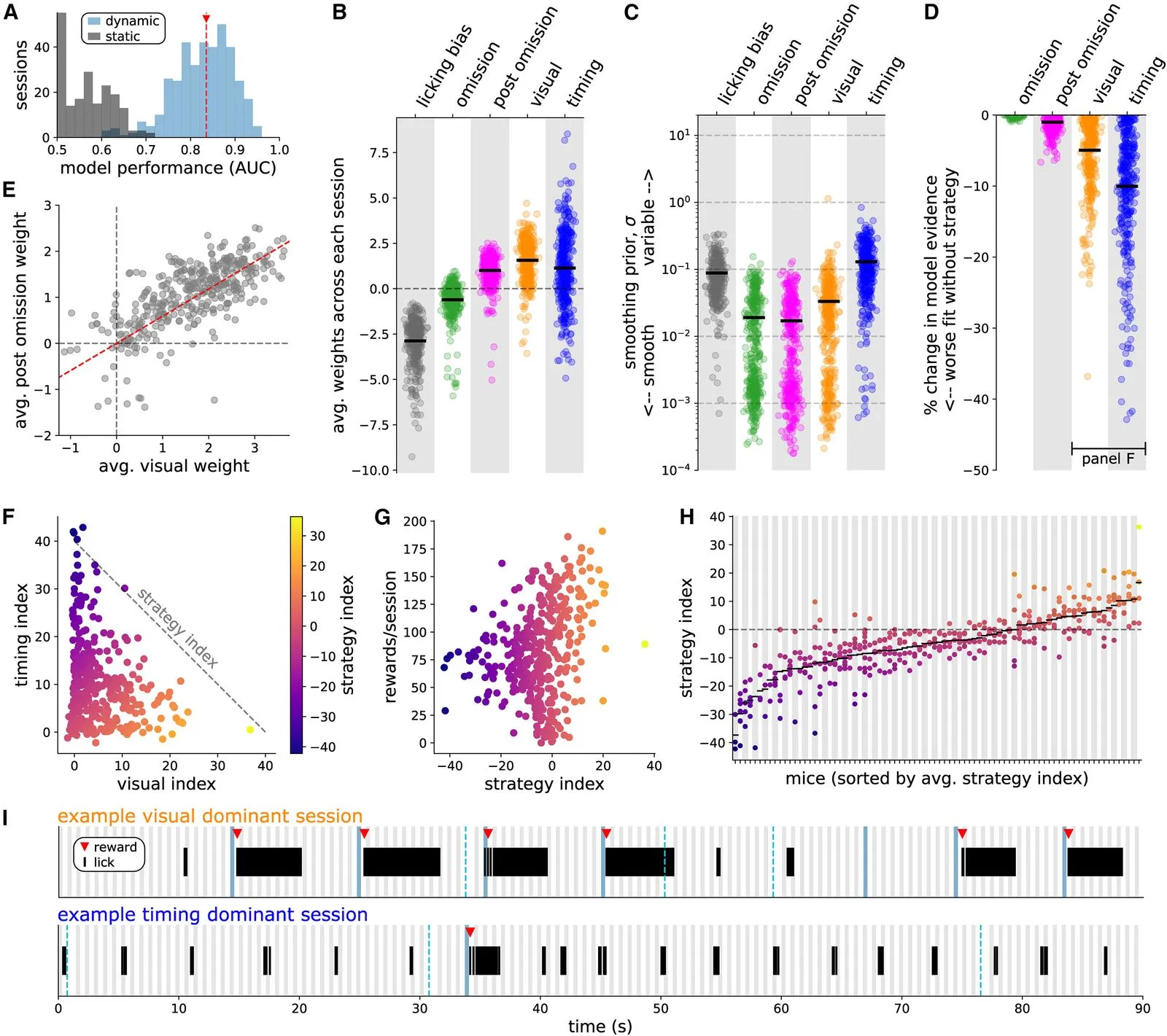
We investigate the question of how distinct animal reward strategies differentially engage brain circuits, focusing on the cortical Vip-Sst disinhibitory circuit between vasoactive intestinal peptide-postive (Vip) interneurons and somatostatin-positive (Sst) interneurons.
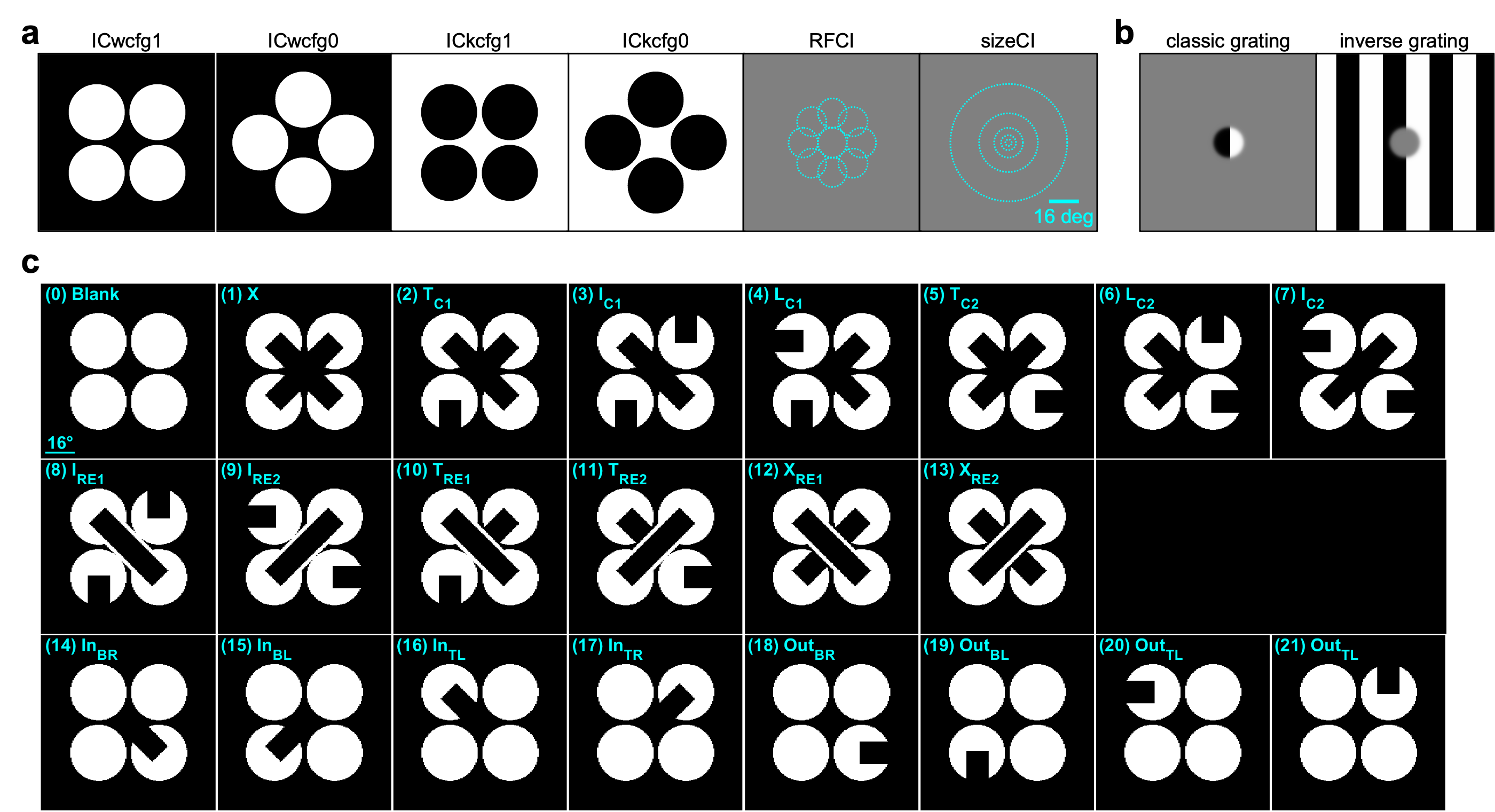

This dataset was collected for the Illusion project, as part of the OpenScope project.
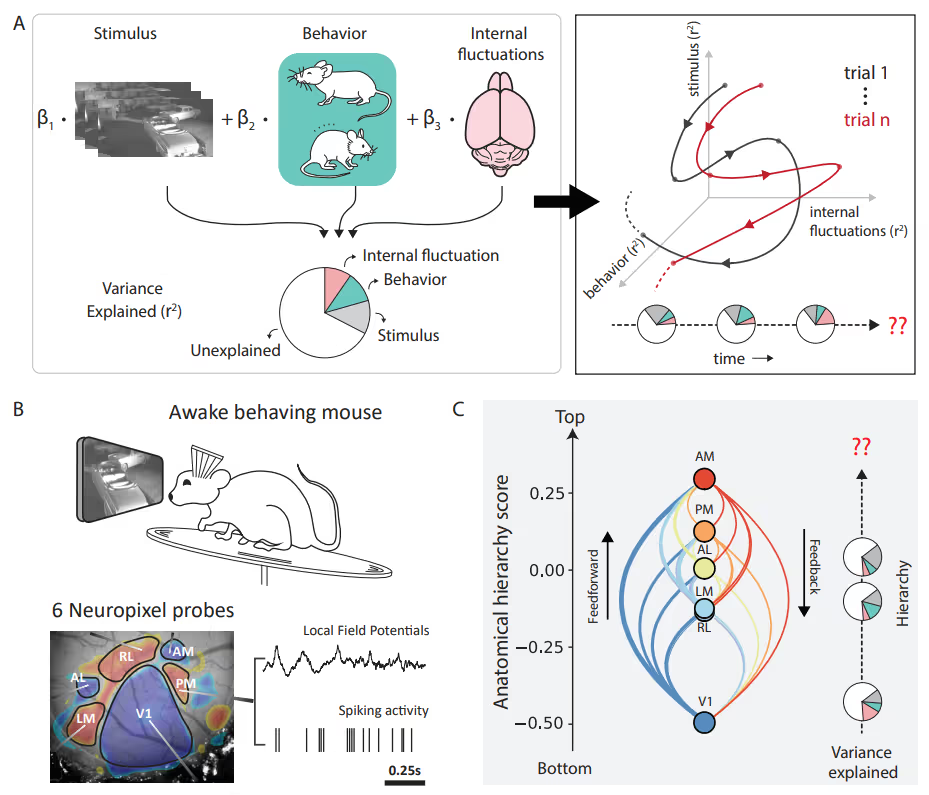
PREPRINT: To address the question of how factors' (such as brain states and behaviors) relative impact neuronal variability evolves over time, we designed an encoding model with latent states to partition visual cortical variability across three crucial categories of sources: internal brain dynamics, behavior, and external visual stimulus.
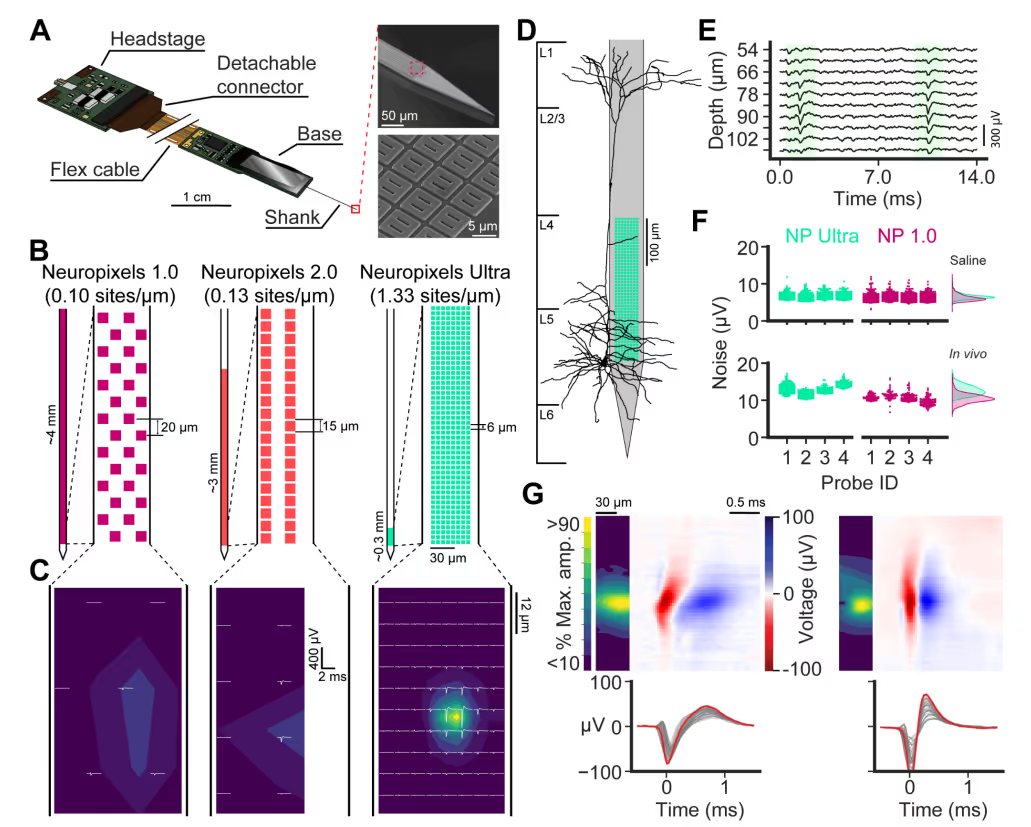
PREPRINT: We developed a silicon probe for extracellular electrophysiology which samples neuronal activity at ultra-high spatial density, allowing us to more accurately and sensitively measure neural activity.
This protocol describes the process of coating an implant with duragel in preparation for in vivo electrophysiology experiments.
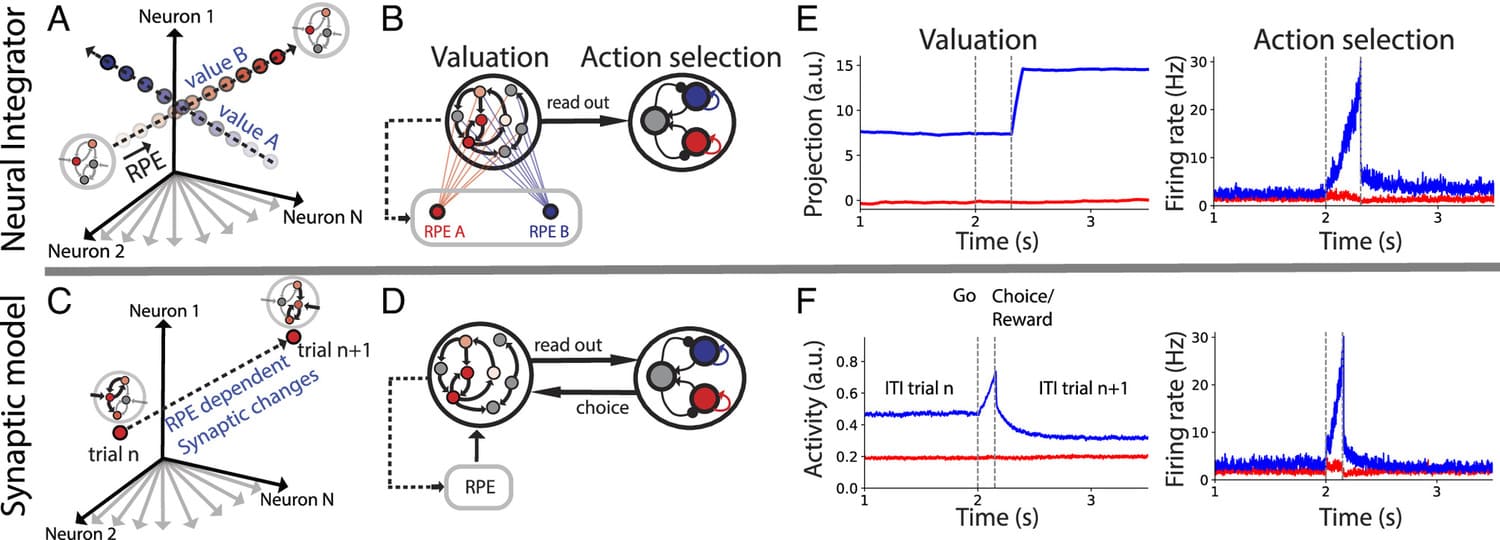
During foraging behavior, action values are persistently encoded in neural activity and updated depending on the history of choice outcomes. What is the neural mechanism for action value maintenance and updating? Here, we explore two contrasting network models: synaptic learning of action value versus neural integration.

This protocol details mounting thin sliced mouse brain tissue sections onto charged slides in a uniform orientation that does not create wrinkles or artifacts that might interfere with imaging, and then how to apply a coverslip.

The Immunohistochemistry (IHC) Staining for Mouse Brain Sections protocol details the blocking, primary, and secondary antibody staining of 50-100 micron mouse brain tissue slices fixed in 4% PFA.

This is a protocol that details sectioning a mouse brain fixed in 4% PFA and 30% sucrose on a sliding microtome.

This protocol collection details the steps for a whole mouse brain specimen to be perfused, sliced, stained with antibodies, DAPI, or both, and made into slides.
This protocol details the steps for staining 4% PFA fixed mouse brain tissue sections with DAPI (4',6-diamidino-2-phenylindole), a fluorescent stain that can be used to label anatomic regions of interest in a mouse brain, allowing for imaging with a fluorescent microscope.
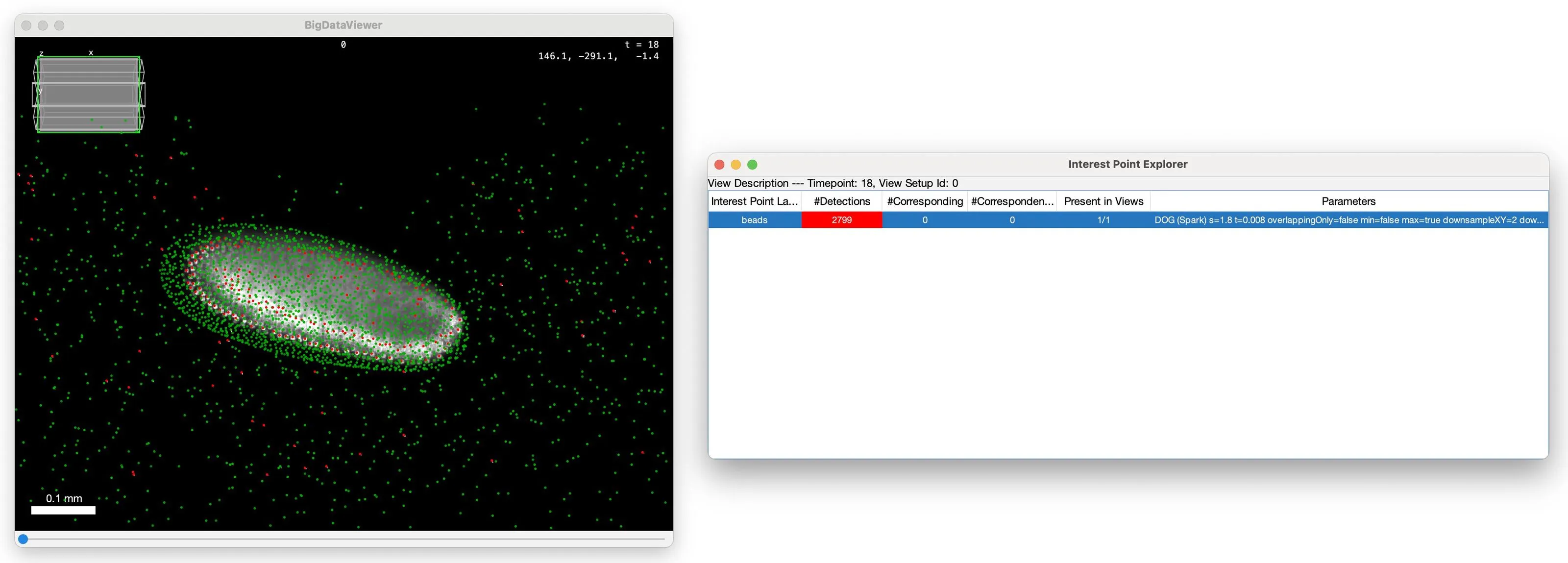
This package allows you to run compute-intense parts of BigStitcher distributed on your workstation, a cluster or the cloud using Apache Spark.
A library for defining and maintaining behavior tasks.
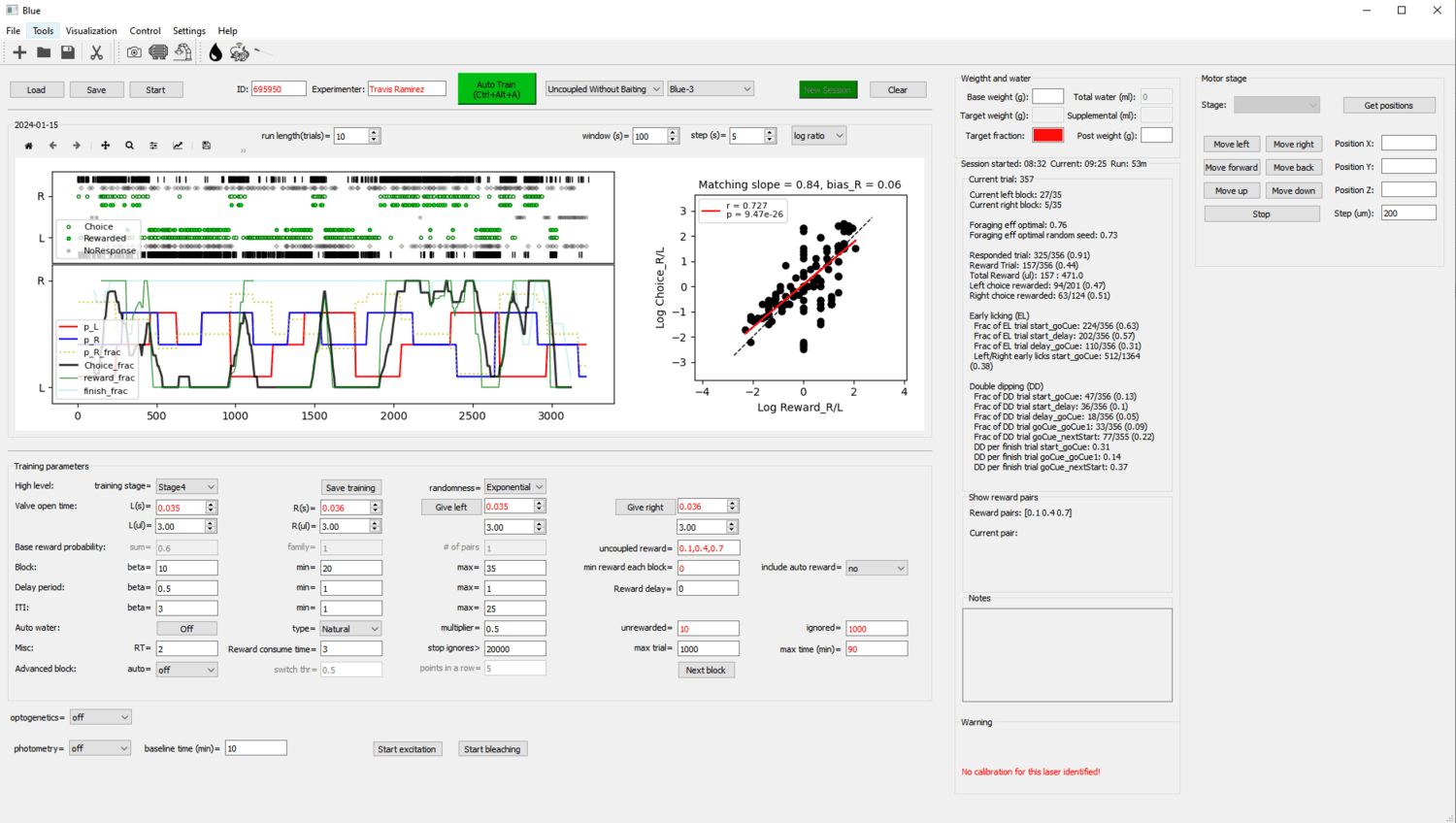
A Bonsai workflow for lick-based foraging experiments, with a PyQt frontend for visualizing performance and modifying task parameters.
This dataset was collected for the Credit Assignment project, as part of the OpenScope project.
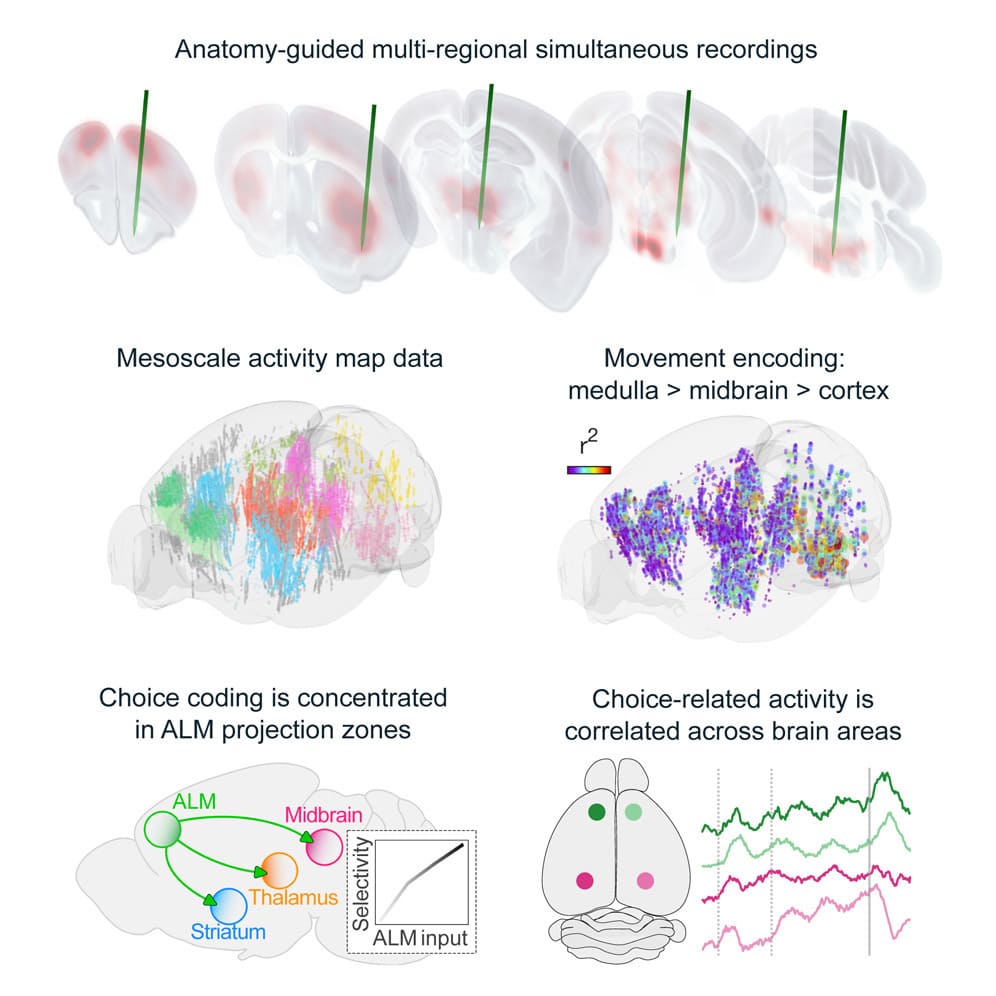
Behavior relies on activity in structured neural circuits that are distributed across the brain, but most experiments probe neurons in a single area at a time. Using multiple Neuropixels probes, we recorded from multi-regional loops connected to the anterior lateral motor cortex (ALM), a circuit node mediating memory-guided directional licking.
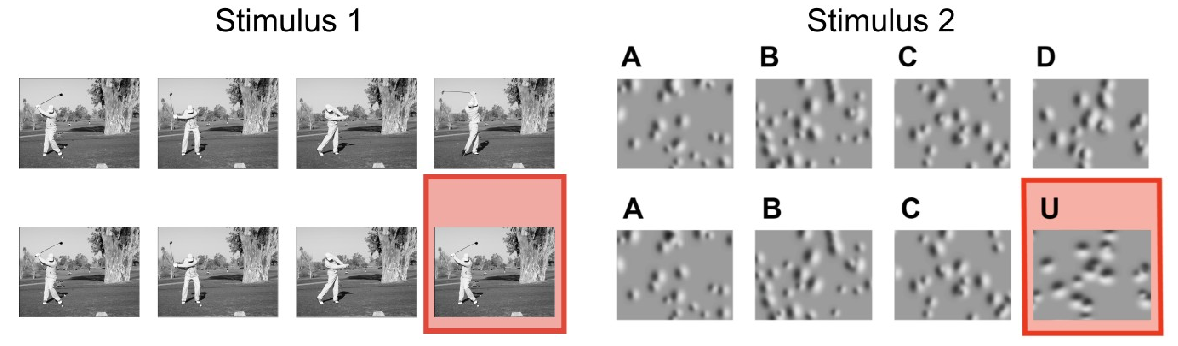

This dataset was collected for the Dendritic Coupling project, as part of the OpenScope project.
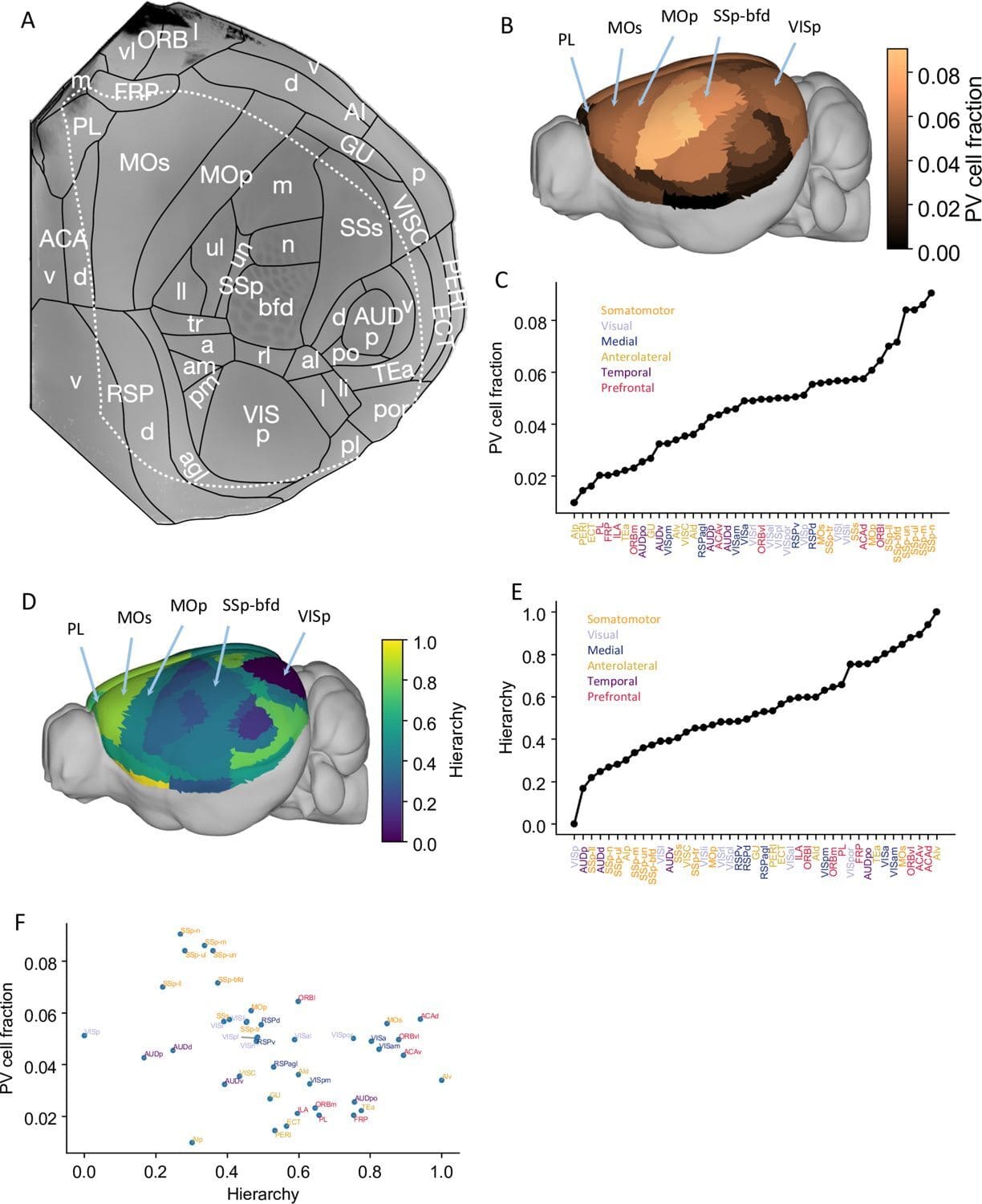
We developed a large-scale model of the multiregional mouse brain for a cardinal cognitive function called working memory, the brain’s ability to internally hold and process information without sensory input.

The OpenScope program publishes on an annual basis a detailed public report that summarizes all of our activities within the year. These reports highlight all developments of the year: new selected projects, completed projects, publications and technical developments of the platform.

TIGRE2-jGCaMP8s-IRES-tTA2-WPRE knock-in mice co-express the jGCaMP8s genetically encoded calcium indicator (GECI) and a tetracycline controlled transactivator (tTA2) in a Cre recombinase-dependent manner from the intergenic Igs7 (TIGRE) locus.

The GENIE Project Team at HHMI Janelia Research Campus have developed and characterized multiple transgenic mice expressing jGCaMP8s and jGCaMP8m.

This protocol provides instructions to prepare the SmartSPIM microscope, to load samples, and to acquire data.
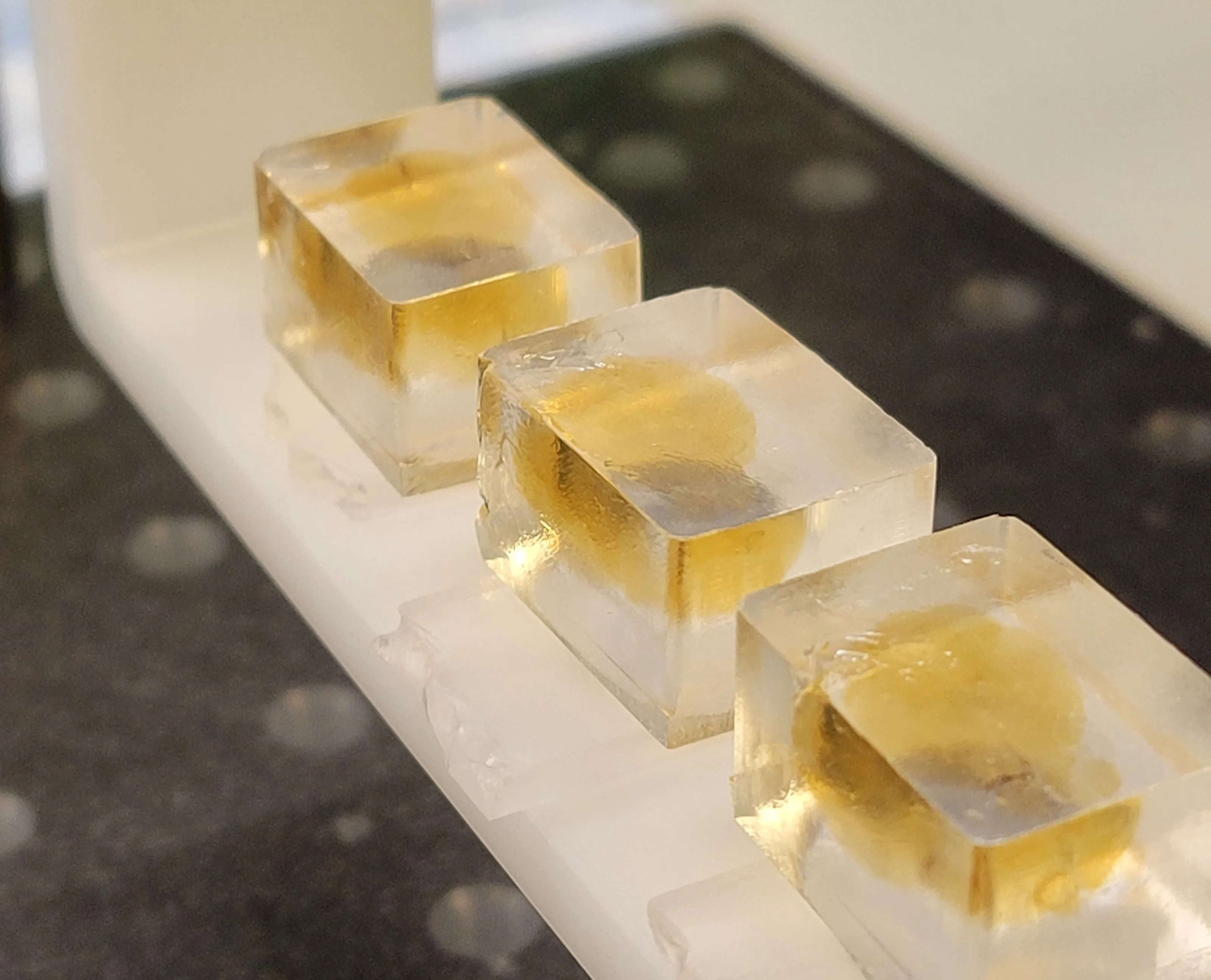

The SmartSPIM is an axially swept lightsheet microscope that produces uniform axial resolution across the entire imaging volume of large biological samples, such as whole mouse brains.
A low-level interface to data collected with the Harp binary protocol.
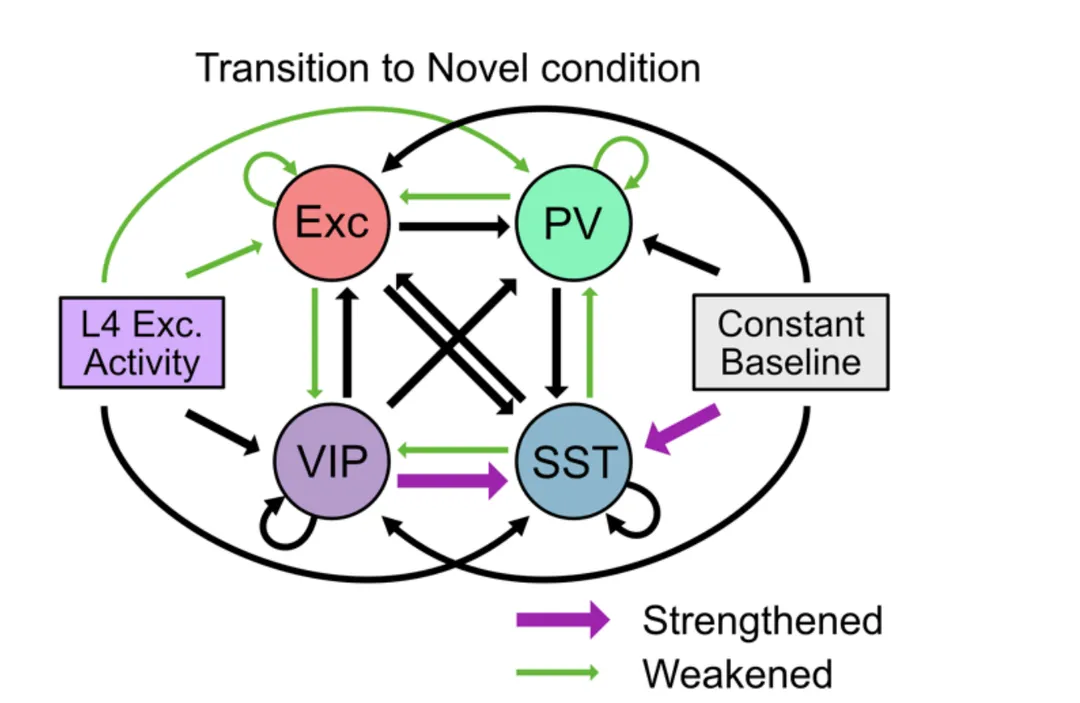
PREPRINT: We employed a large-scale public 16 dataset of electrophysiological recordings in the visual cortex of awake, behaving mice using 17 Neuropixels probes and designed population network models to investigate the observed 18 changes in neural dynamics in response to a combination of distinct forms of novelty.
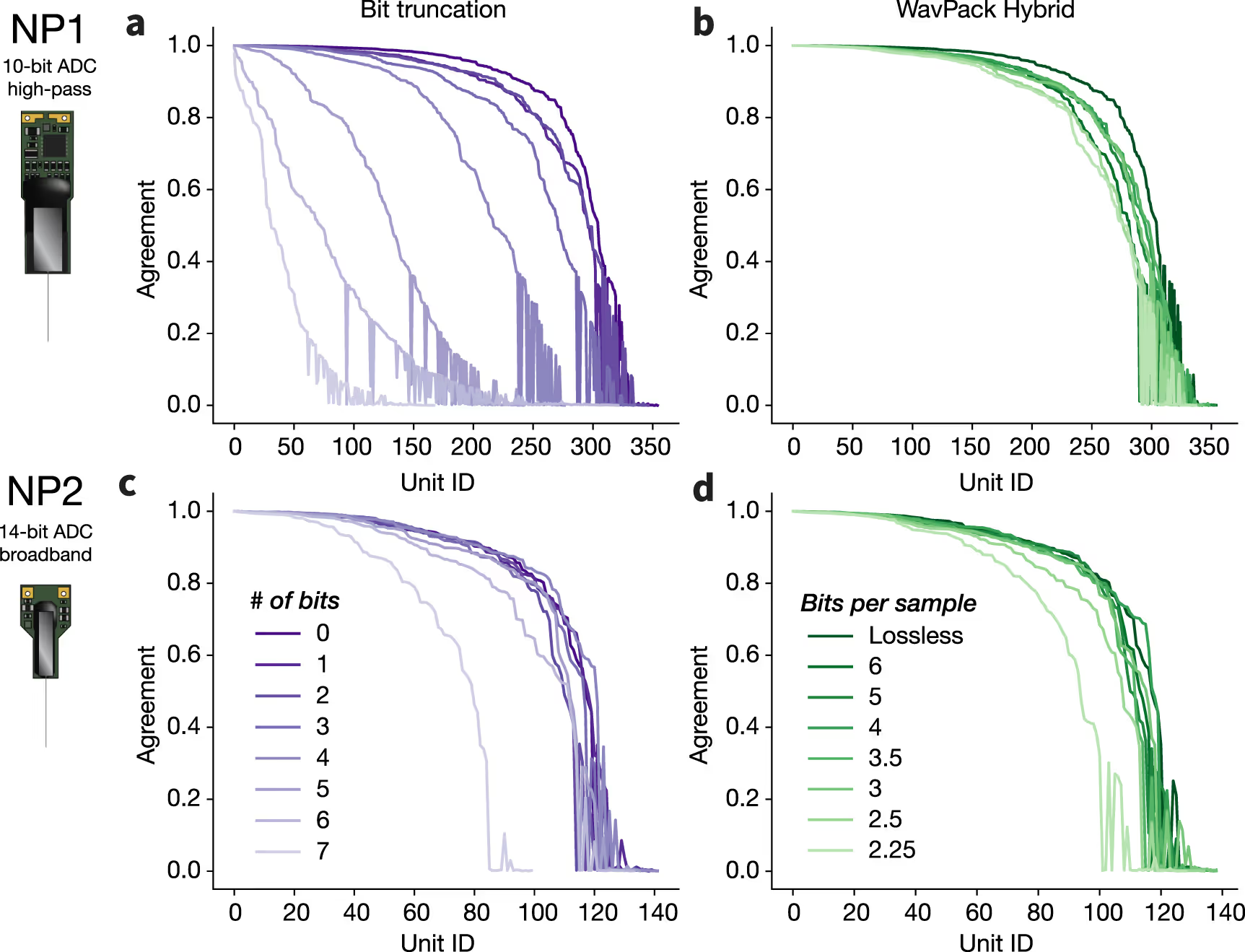
We establish a set of benchmarks for comparing the performance of various compression algorithms on experimental and simulated recordings from Neuropixels 1.0 (NP1) and 2.0 (NP2) probes.
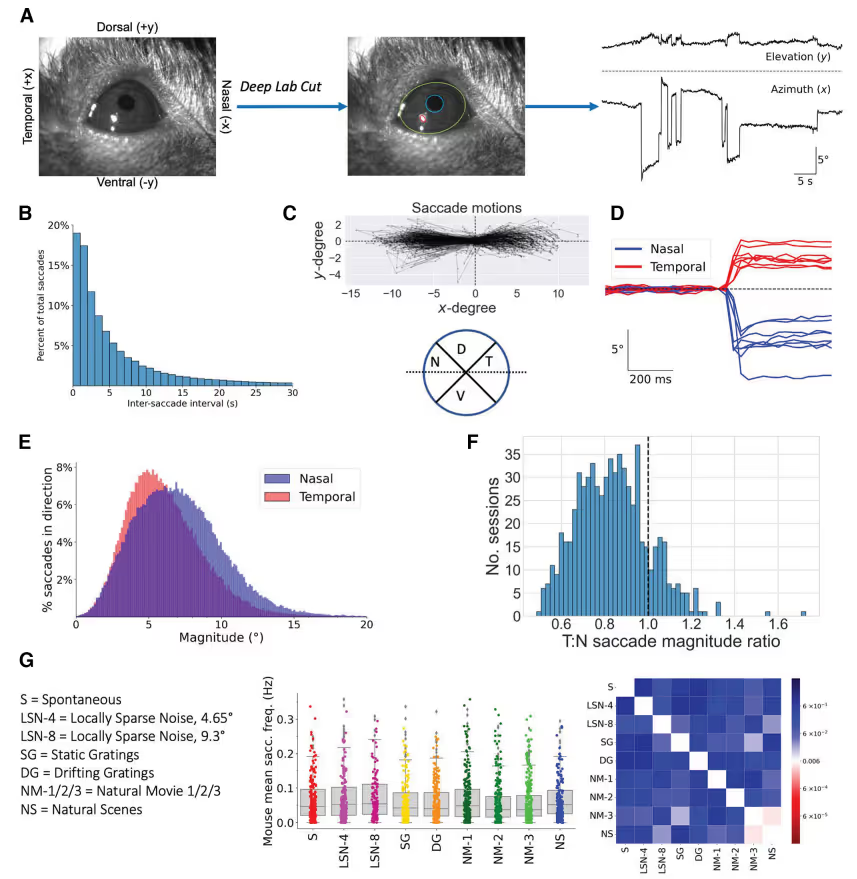
We present a large-scale analysis of saccadic behaviors in head-fixed mice and their neural correlates. We find that saccade-responsive neurons are present across visual cortex, but their distribution varies considerably by transgenically defined cell type, cortical area, and cortical layer.

Facilitating Neuronal Structure and Function Analysis: Learn about VAST's pivotal role in supporting the Allen Institute's data-intensive machine learning workloads, facilitating their analysis of neuronal structure and function.
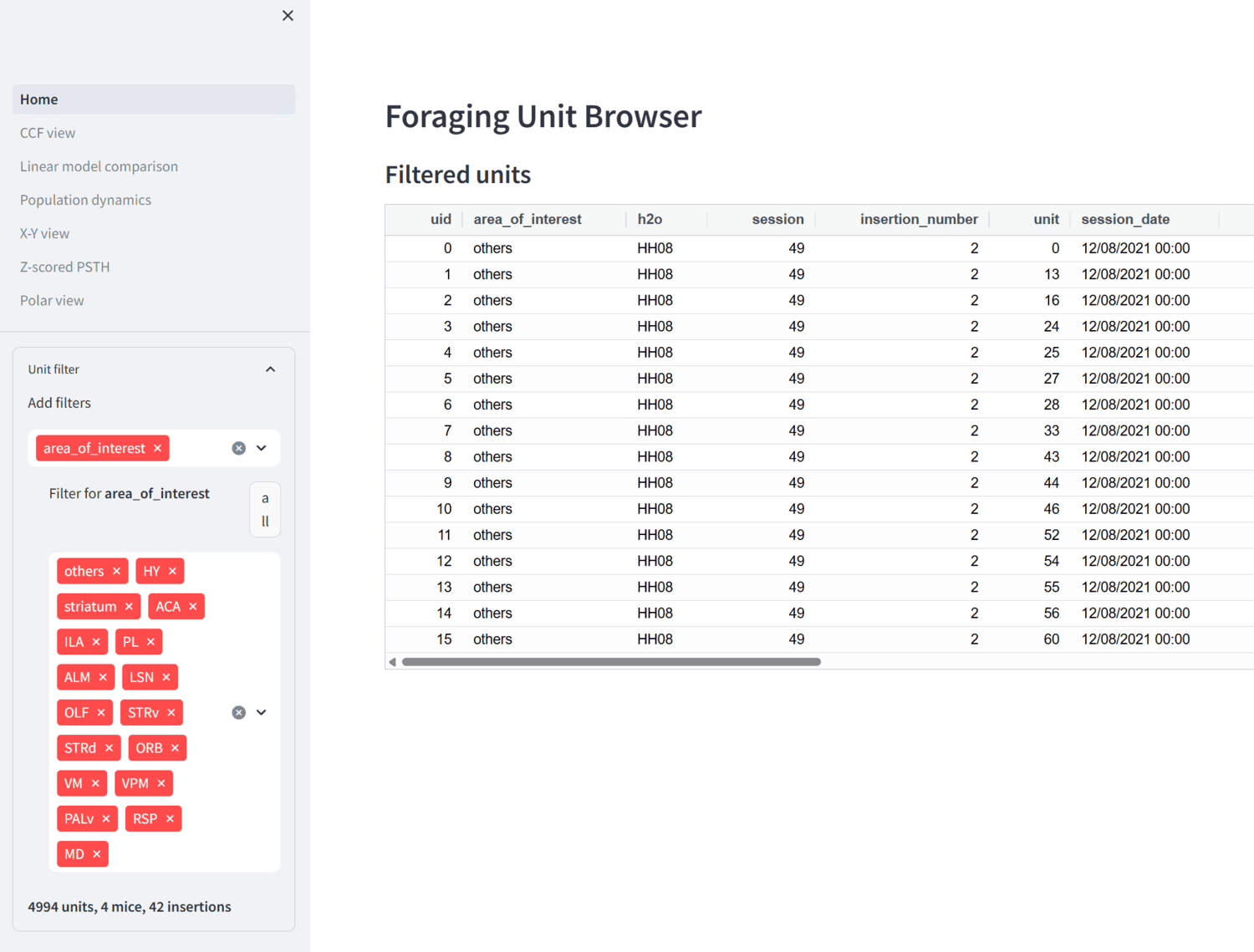
Streamlit app for browsing foraging ephys data.
Using technology originally designed for defect detection in electronics manufacturing, the newly built “ExA-SPIM” microscope is showing scientists the mouse brain as it’s never been seen before.

PREPRINT: Using cellular resolution, mesoscale two-photon calcium imaging and multi-Neuropixels recordings in the mouse visual cortex, we identified a sparse subset of neurons in the primary visual cortex (V1) and higher visual areas that respond emergently to illusory contours, which are key tools to study sensory inference.
We have imaged expanded whole mouse brains generated using this protocol on our custom built ExA-SPIM microscope without need for any tissue slicing. These whole brain data sets are being used for tracing complete axonal morphology of individual neurons.
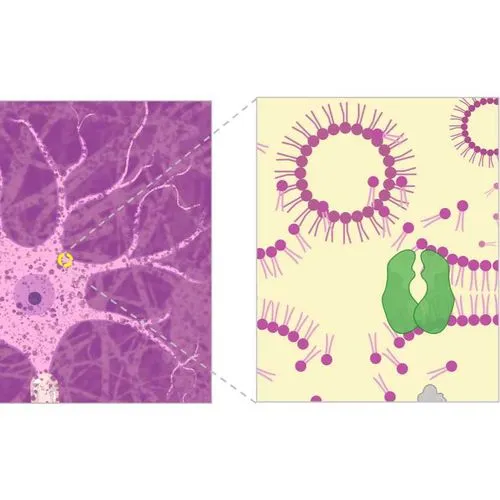
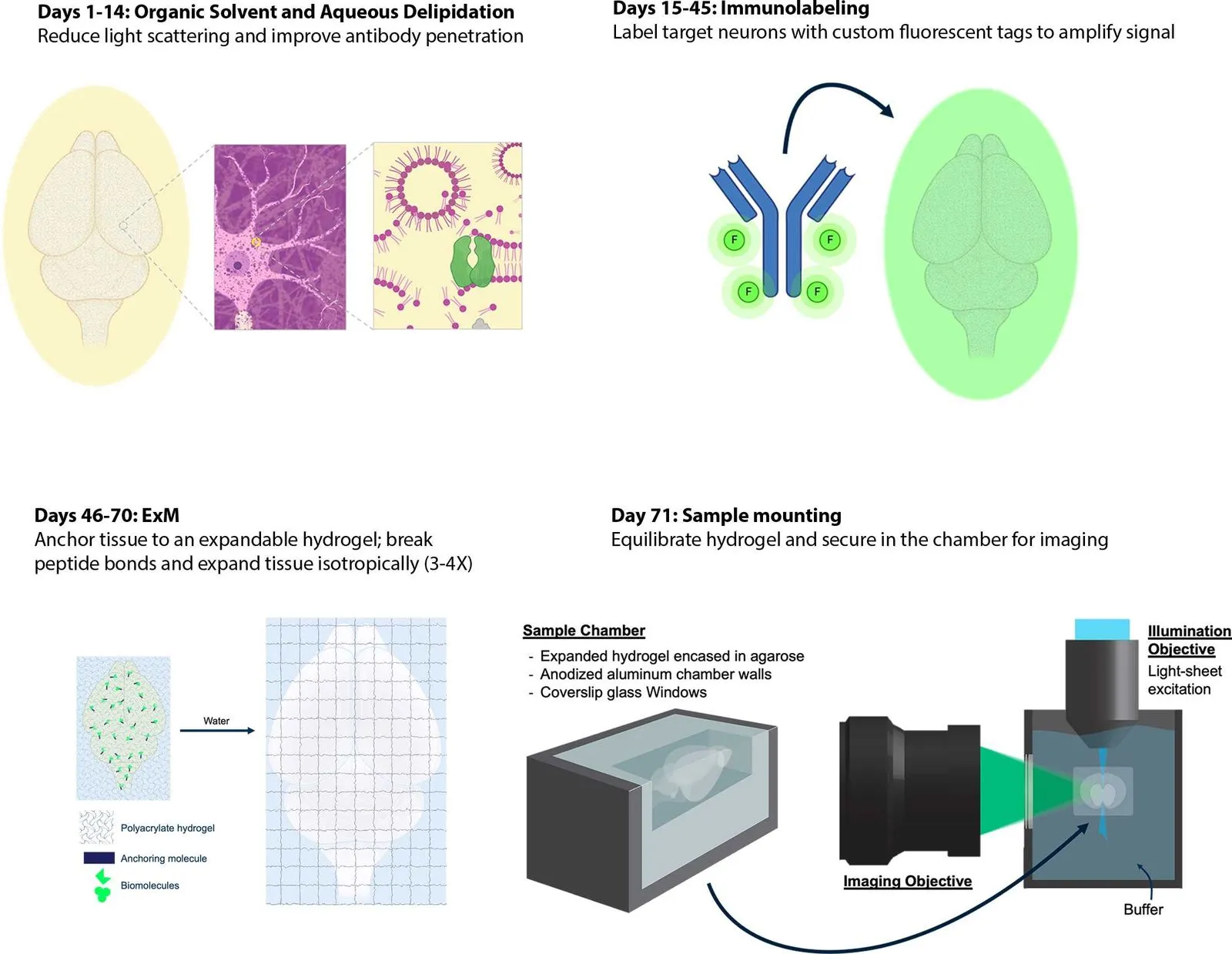
When paired with organic delipidation, these steps can return a solvent-shrunken brain to normal size in phosphate buffer, suitable for post-delipidation antibody labeling.

To amplify signals of FPs for high resolution, multi-scale imaging, we immunolabel delipidated brain samples.

This protocol produces a delipidated whole brain that can be further processed for expansion or index-matched for immediate imaging.
This dataset was collected for the Predictive Coding project, as part of the OpenScope project.

This repository contains CAD (Solidworks) files for the ExA-SPIM microscope system.
This repository contains optical (ZEMAX) files for the ExA-SPIM microscope system.
This study investigates the effects of periodic sensory stimulation (PSS) on the activity of neurons of different types in the mouse visual cortex.
We used two-photon calcium imaging in mice of both sexes to systematically survey stimulus-evoked neurophysiological differentiation (ND) in excitatory neuronal populations in layers (L)2/3, L4, and L5 across five visual cortical areas (primary, lateromedial, anterolateral, posteromedial, and anteromedial) in response to naturalistic and phase-scrambled movie stimuli.
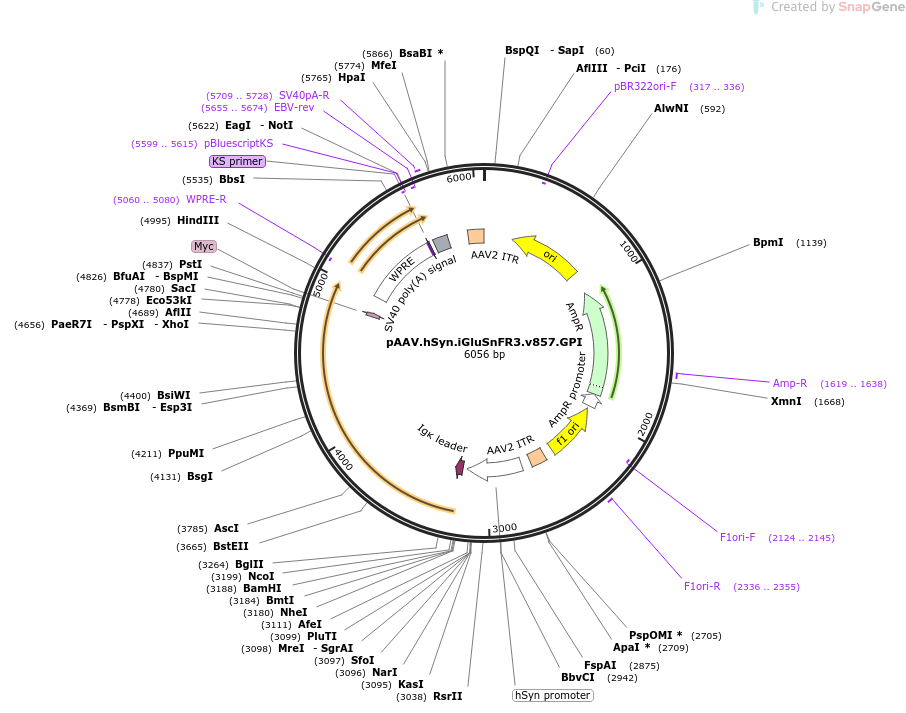
Fluorescent reporters for glutamate, third generation, variant 857. iGluSnFR3 exhibits rapid non-saturating activation kinetics and reports synaptic glutamate release with decreased saturation and increased specificity versus extra-synaptic signals in cultured neurons.

We generated variants of the fluorescent glutamate indicator iGluSnFR with improved signal-to-noise ratios and kinetics, to enable better imaging of neurotransmission with genetic and molecular specificity.
Code Ocean capsule for sharing analysis of thalamus MERFISH data, primarily in jupyter notebook form.
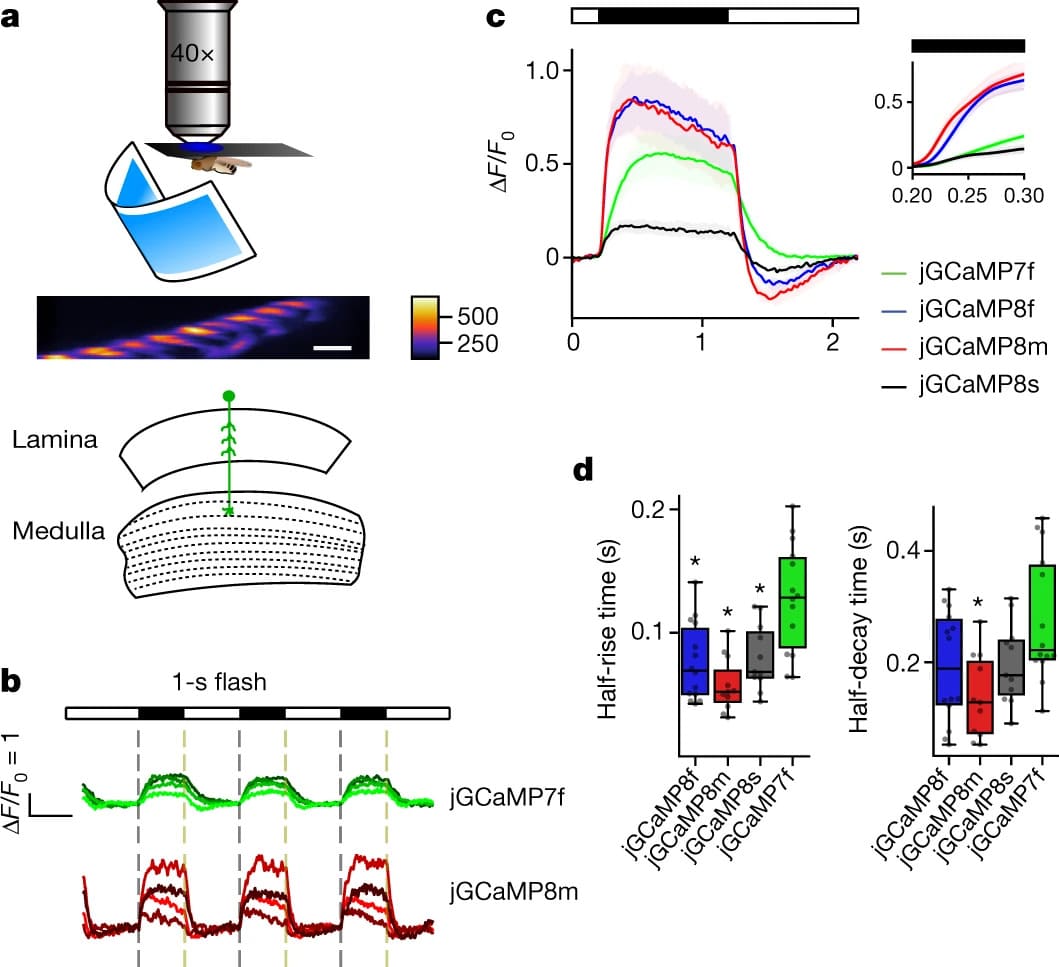
Here we used large-scale screening and structure-guided mutagenesis to develop and optimize several fast and sensitive GCaMP-type indicators. jGCaMP8 sensors will allow tracking of large populations of neurons on timescales relevant to neural computation.
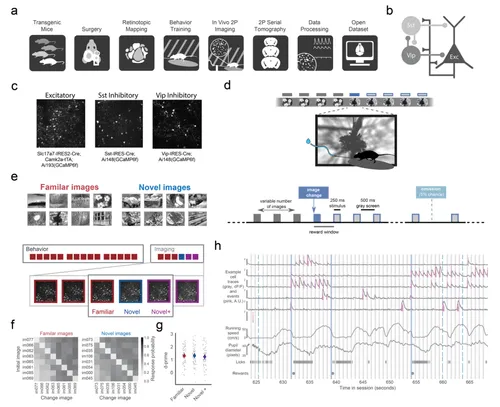
PREPRINT: We characterize the impact of stimulus novelty on disinhibitory circuit components using longitudinal 2-photon calcium imaging of Vip, Sst, and excitatory populations in the mouse visual cortex.

Workspace for AIND protocols.

The OpenScope program publishes on an annual basis a detailed public report that summarizes all of our activities within the year. These reports highlight all developments of the year: new selected projects, completed projects, publications and technical developments of the platform.

The OpenScope program publishes on an annual basis a detailed public report that summarizes all of our activities within the year. These reports highlight all developments of the year: new selected projects, completed projects, publications and technical developments of the platform.

Here we report the engineering strategies that we implemented when creating the Allen Brain Observatory physiology pipelines.

Selected design files for custom hardware for scientific instruments that we have developed for use in-house, or in partnership with collaborators.

This protocol describes the delivery of a neuronal tracer using the iontophoretic method. The surgery uses a stereotaxic system to target specific brain coordinates in the mouse.

This protocol describes the delivery of a neuronal tracer using the Nanoject II. The surgery uses a stereotaxic system to target specific brain coordinates in the mouse.
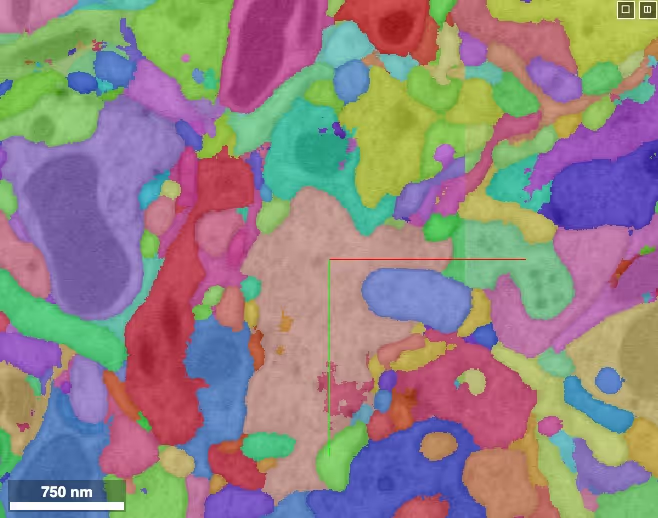
Set of tools that work with and enhance the Google neuroglancer tool for volume visualization and annotation.
SpikeInterface is a Python framework designed to unify preexisting spike sorting technologies into a single code base.

This webpage stores metadata, experimental details and project description associated and ran by the OpenScope program. OpenScope is a platform for high-throughput and reproducible neurophysiology open to external scientists to test theories of brain function.

Thorlabs' 2-Photon Random Access Mesoscope provides subcellular resolution over an exceptionally large 5 mm x 5 mm field of view.
Neuroglancer is a WebGL-based viewer for volumetric data. It is capable of displaying arbitrary (non axis-aligned) cross-sectional views of volumetric data, as well as 3-D meshes and line-segment based models (skeletons).
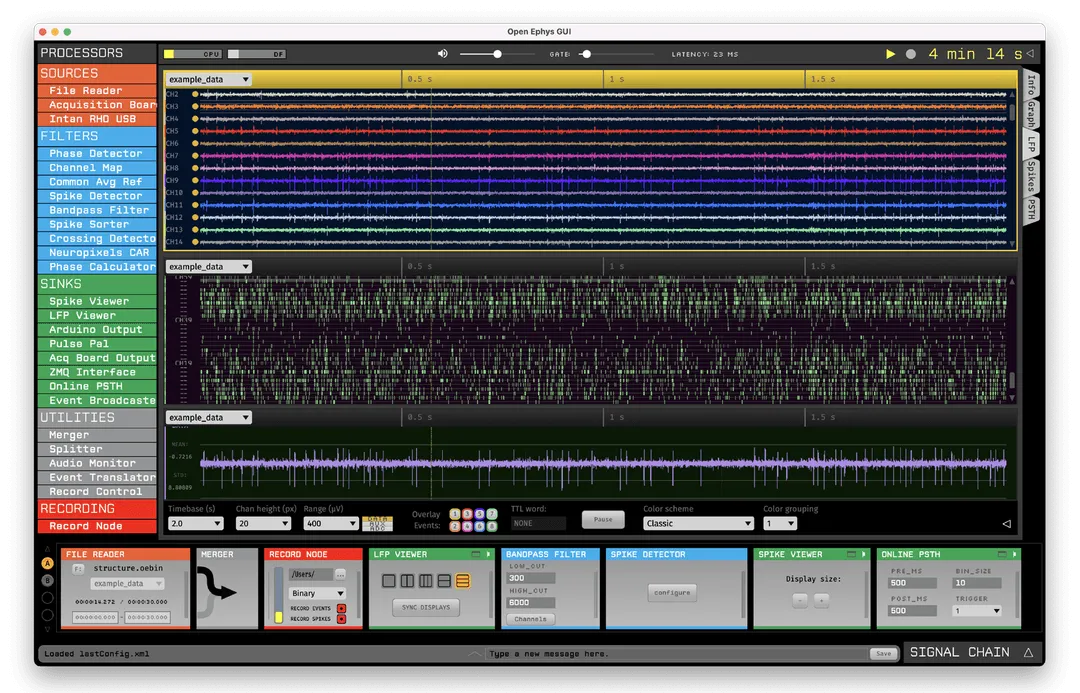
The Open Ephys GUI was built by neuroscientists, for neuroscientists. It has all the features needed to acquire and visualize your data—while also making it easy to add new modules written in C++.
%20(1).jpg)
We study the biological mechanisms that enable animals to learn the structure of their world through free exploration. To do this, we collect long-term, uninterrupted records of the natural behavior, brain activity, and network connectivity of mice while they repeatedly interact with odors in their environment.
.png)
We are developing and using genetic, electrophysiological, optical, and behavioral approaches to investigate how the brain adaptively controls behavior. The team focuses on understanding the descending circuits that control the execution of actions and how they change when actions are reinforced and refined.
We are using large-scale electrophysiology to study how distributed brain regions coordinate their spiking activity to guide behavior in changing environments.